|
Symbol Name ID |
Cybb
cytochrome b-245, beta polypeptide MGI:88574 |
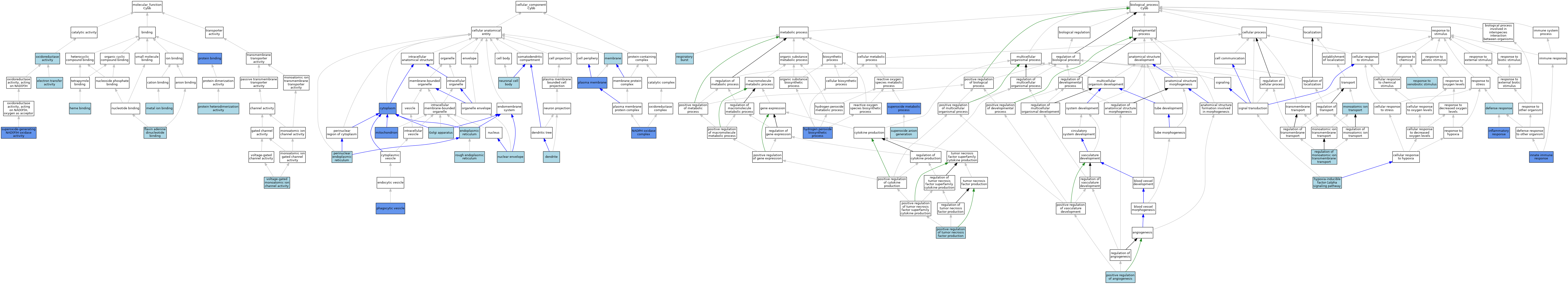
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0009055 | electron transfer activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0020037 | heme binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0046872 | metal ion binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016491 | oxidoreductase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016491 | oxidoreductase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:172364 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:209569 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:209569 | |||||||||
| Molecular Function | GO:0046982 | protein heterodimerization activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0016175 | superoxide-generating NAD(P)H oxidase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0016175 | superoxide-generating NAD(P)H oxidase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0016175 | superoxide-generating NAD(P)H oxidase activity | IDA | J:185616 | |||||||||
| Molecular Function | GO:0005244 | voltage-gated monoatomic ion channel activity | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:193251 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:103853 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:103853 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:97972 | |||||||||
| Cellular Component | GO:0030425 | dendrite | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005783 | endoplasmic reticulum | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005794 | Golgi apparatus | ISO | J:155856 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:72247 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0043020 | NADPH oxidase complex | ISO | J:155856 | |||||||||
| Cellular Component | GO:0043020 | NADPH oxidase complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0043020 | NADPH oxidase complex | IDA | J:209569 | |||||||||
| Cellular Component | GO:0043020 | NADPH oxidase complex | IBA | J:265628 | |||||||||
| Cellular Component | GO:0043025 | neuronal cell body | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005635 | nuclear envelope | ISO | J:155856 | |||||||||
| Cellular Component | GO:0097038 | perinuclear endoplasmic reticulum | ISO | J:155856 | |||||||||
| Cellular Component | GO:0045335 | phagocytic vesicle | IDA | J:193251 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IDA | J:103853 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IDA | J:103853 | |||||||||
| Cellular Component | GO:0005791 | rough endoplasmic reticulum | ISO | J:155856 | |||||||||
| Biological Process | GO:0006952 | defense response | IBA | J:265628 | |||||||||
| Biological Process | GO:0050665 | hydrogen peroxide biosynthetic process | IMP | J:91766 | |||||||||
| Biological Process | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway | ISO | J:155856 | |||||||||
| Biological Process | GO:0006954 | inflammatory response | IMP | J:209569 | |||||||||
| Biological Process | GO:0045087 | innate immune response | IMP | J:241355 | |||||||||
| Biological Process | GO:0045087 | innate immune response | ISO | J:164563 | |||||||||
| Biological Process | GO:0006811 | monoatomic ion transport | IEA | J:60000 | |||||||||
| Biological Process | GO:0045766 | positive regulation of angiogenesis | ISO | J:155856 | |||||||||
| Biological Process | GO:0032760 | positive regulation of tumor necrosis factor production | ISO | J:155856 | |||||||||
| Biological Process | GO:0034765 | regulation of monoatomic ion transmembrane transport | IEA | J:60000 | |||||||||
| Biological Process | GO:0045730 | respiratory burst | ISO | J:164563 | |||||||||
| Biological Process | GO:0009410 | response to xenobiotic stimulus | ISO | J:155856 | |||||||||
| Biological Process | GO:0042554 | superoxide anion generation | ISO | J:164563 | |||||||||
| Biological Process | GO:0042554 | superoxide anion generation | IBA | J:265628 | |||||||||
| Biological Process | GO:0006801 | superoxide metabolic process | IMP | J:209569 | |||||||||
| Biological Process | GO:0006801 | superoxide metabolic process | ISO | J:164563 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||