|
Symbol Name ID |
Nlrp10
NLR family, pyrin domain containing 10 MGI:2444084 |
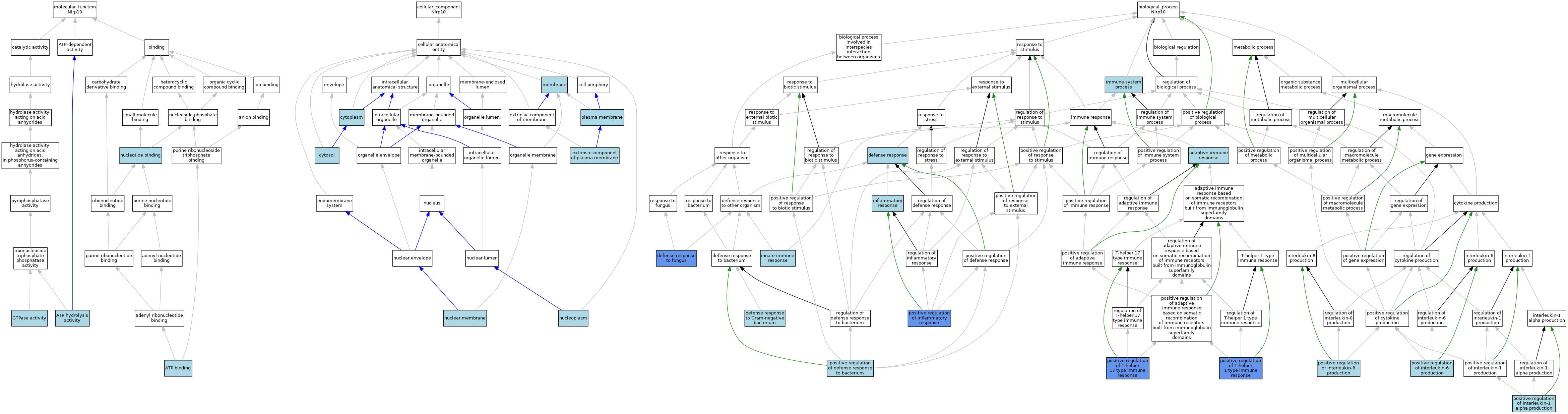
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016887 | ATP hydrolysis activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0003924 | GTPase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0019897 | extrinsic component of plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0031965 | nuclear membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IEA | J:60000 | |||||||||
| Biological Process | GO:0002250 | adaptive immune response | IEA | J:60000 | |||||||||
| Biological Process | GO:0006952 | defense response | RCA | J:84168 | |||||||||
| Biological Process | GO:0050832 | defense response to fungus | IMP | J:190699 | |||||||||
| Biological Process | GO:0050829 | defense response to Gram-negative bacterium | ISO | J:164563 | |||||||||
| Biological Process | GO:0002376 | immune system process | IEA | J:60000 | |||||||||
| Biological Process | GO:0006954 | inflammatory response | IEA | J:60000 | |||||||||
| Biological Process | GO:0045087 | innate immune response | IEA | J:60000 | |||||||||
| Biological Process | GO:1900426 | positive regulation of defense response to bacterium | ISO | J:164563 | |||||||||
| Biological Process | GO:0050729 | positive regulation of inflammatory response | IMP | J:245258 | |||||||||
| Biological Process | GO:0050729 | positive regulation of inflammatory response | IBA | J:265628 | |||||||||
| Biological Process | GO:0032730 | positive regulation of interleukin-1 alpha production | ISO | J:164563 | |||||||||
| Biological Process | GO:0032755 | positive regulation of interleukin-6 production | ISO | J:164563 | |||||||||
| Biological Process | GO:0032757 | positive regulation of interleukin-8 production | ISO | J:164563 | |||||||||
| Biological Process | GO:0002827 | positive regulation of T-helper 1 type immune response | IMP | J:190699 | |||||||||
| Biological Process | GO:2000318 | positive regulation of T-helper 17 type immune response | IMP | J:190699 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||