|
Symbol Name ID |
Atxn7
ataxin 7 MGI:2179277 |
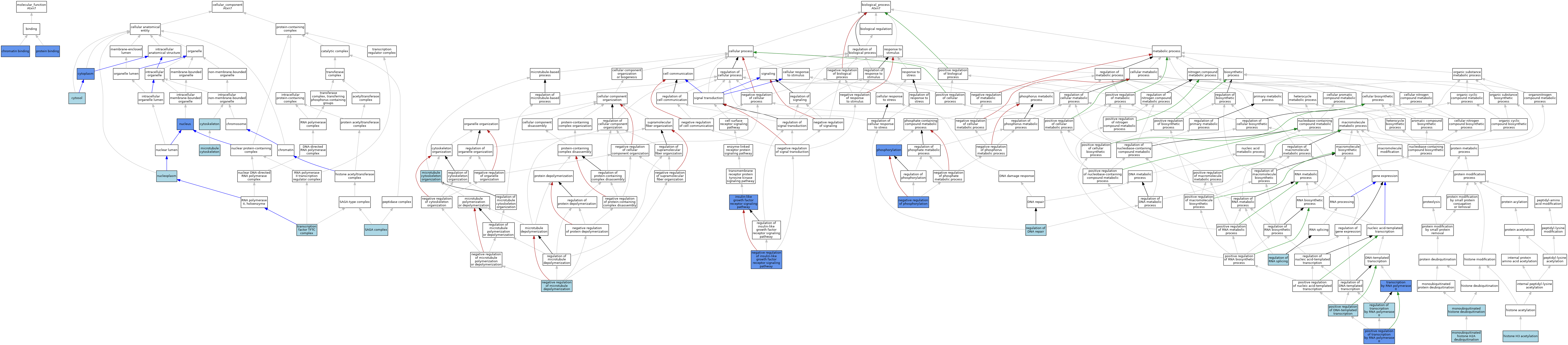
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0003682 | chromatin binding | IDA | J:87337 | |||||||||
| Molecular Function | GO:0003682 | chromatin binding | IDA | J:87337 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:107098 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:162104 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:87337 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:75988 | |||||||||
| Cellular Component | GO:0005856 | cytoskeleton | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0015630 | microtubule cytoskeleton | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IDA | J:75988 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IDA | J:87337 | |||||||||
| Cellular Component | GO:0000124 | SAGA complex | NAS | J:320037 | |||||||||
| Cellular Component | GO:0033276 | transcription factor TFTC complex | NAS | J:320038 | |||||||||
| Biological Process | GO:0043966 | histone H3 acetylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0043966 | histone H3 acetylation | NAS | J:320036 | |||||||||
| Biological Process | GO:0048009 | insulin-like growth factor receptor signaling pathway | IMP | J:131821 | |||||||||
| Biological Process | GO:0048009 | insulin-like growth factor receptor signaling pathway | IMP | J:131821 | |||||||||
| Biological Process | GO:0000226 | microtubule cytoskeleton organization | ISO | J:164563 | |||||||||
| Biological Process | GO:0035521 | monoubiquitinated histone deubiquitination | ISO | J:164563 | |||||||||
| Biological Process | GO:0035522 | monoubiquitinated histone H2A deubiquitination | ISO | J:164563 | |||||||||
| Biological Process | GO:0043569 | negative regulation of insulin-like growth factor receptor signaling pathway | IMP | J:131821 | |||||||||
| Biological Process | GO:0007026 | negative regulation of microtubule depolymerization | ISO | J:164563 | |||||||||
| Biological Process | GO:0042326 | negative regulation of phosphorylation | IMP | J:131821 | |||||||||
| Biological Process | GO:0016310 | phosphorylation | IMP | J:131821 | |||||||||
| Biological Process | GO:0016310 | phosphorylation | IMP | J:131821 | |||||||||
| Biological Process | GO:0045893 | positive regulation of DNA-templated transcription | NAS | J:320111 | |||||||||
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | IMP | J:131821 | |||||||||
| Biological Process | GO:0006282 | regulation of DNA repair | NAS | J:320036 | |||||||||
| Biological Process | GO:0006282 | regulation of DNA repair | NAS | J:320001 | |||||||||
| Biological Process | GO:0043484 | regulation of RNA splicing | NAS | J:320036 | |||||||||
| Biological Process | GO:0006357 | regulation of transcription by RNA polymerase II | ISO | J:164563 | |||||||||
| Biological Process | GO:0006366 | transcription by RNA polymerase II | IMP | J:131821 | |||||||||
| Biological Process | GO:0006366 | transcription by RNA polymerase II | IMP | J:131821 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||