|
Symbol Name ID |
Smc1b
structural maintenance of chromosomes 1B MGI:2154049 |
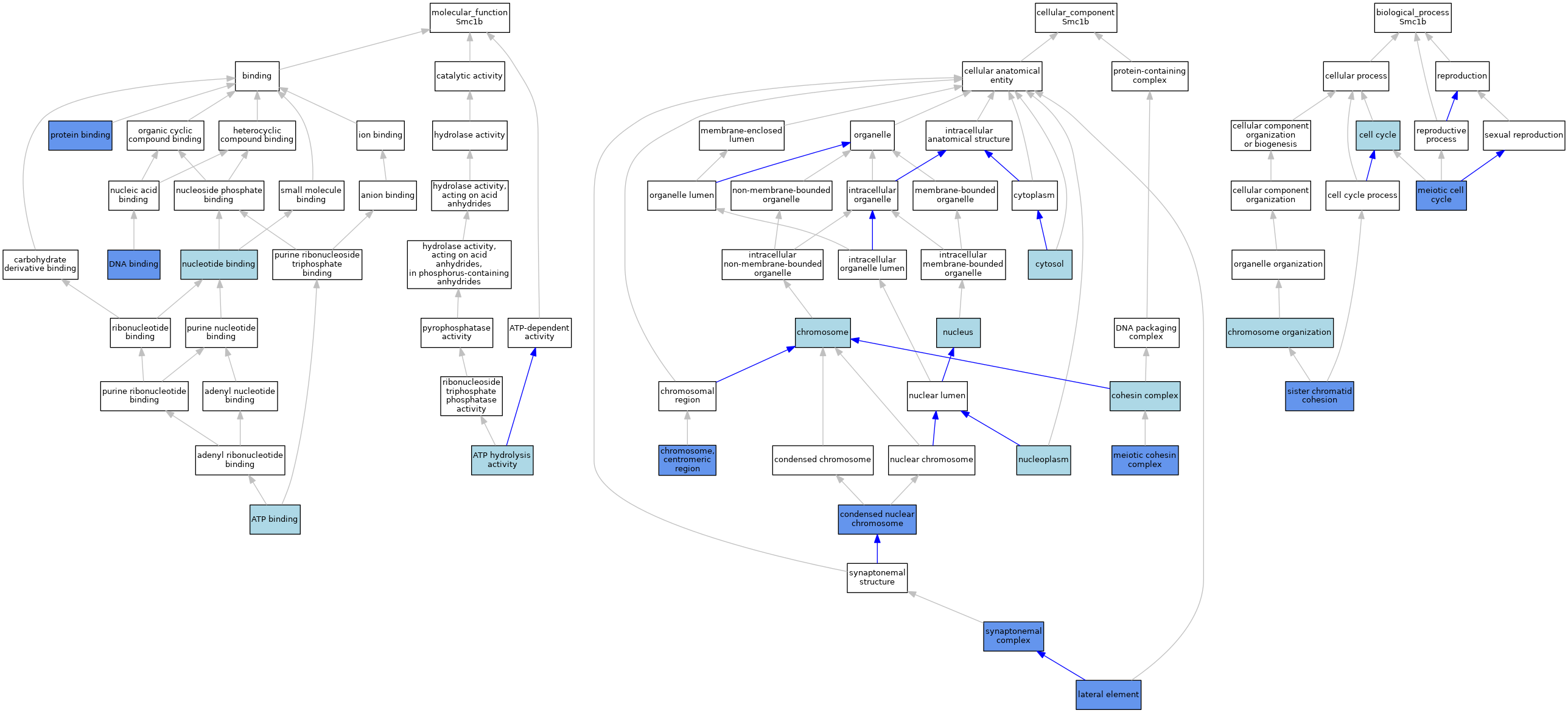
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016887 | ATP hydrolysis activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0003677 | DNA binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0003677 | DNA binding | IDA | J:71579 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:71579 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:84400 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:84400 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:84400 | |||||||||
| Cellular Component | GO:0005694 | chromosome | IEA | J:72247 | |||||||||
| Cellular Component | GO:0005694 | chromosome | IEA | J:60000 | |||||||||
| Cellular Component | GO:0000775 | chromosome, centromeric region | IDA | J:71579 | |||||||||
| Cellular Component | GO:0008278 | cohesin complex | IBA | J:265628 | |||||||||
| Cellular Component | GO:0000794 | condensed nuclear chromosome | IDA | J:182264 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0000800 | lateral element | IDA | J:98372 | |||||||||
| Cellular Component | GO:0000800 | lateral element | IDA | J:185266 | |||||||||
| Cellular Component | GO:0030893 | meiotic cohesin complex | IDA | J:173689 | |||||||||
| Cellular Component | GO:0030893 | meiotic cohesin complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0030893 | meiotic cohesin complex | IDA | J:176168 | |||||||||
| Cellular Component | GO:0030893 | meiotic cohesin complex | IDA | J:182264 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912355 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912362 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912423 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912449 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912468 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912506 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IBA | J:265628 | |||||||||
| Cellular Component | GO:0000795 | synaptonemal complex | IDA | J:71579 | |||||||||
| Cellular Component | GO:0000795 | synaptonemal complex | IDA | J:113188 | |||||||||
| Biological Process | GO:0007049 | cell cycle | IEA | J:60000 | |||||||||
| Biological Process | GO:0051276 | chromosome organization | IEA | J:72247 | |||||||||
| Biological Process | GO:0051321 | meiotic cell cycle | IPI | J:84400 | |||||||||
| Biological Process | GO:0007062 | sister chromatid cohesion | IBA | J:265628 | |||||||||
| Biological Process | GO:0007062 | sister chromatid cohesion | IDA | J:71579 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||