|
Symbol Name ID |
Dlst
dihydrolipoamide S-succinyltransferase MGI:1926170 |
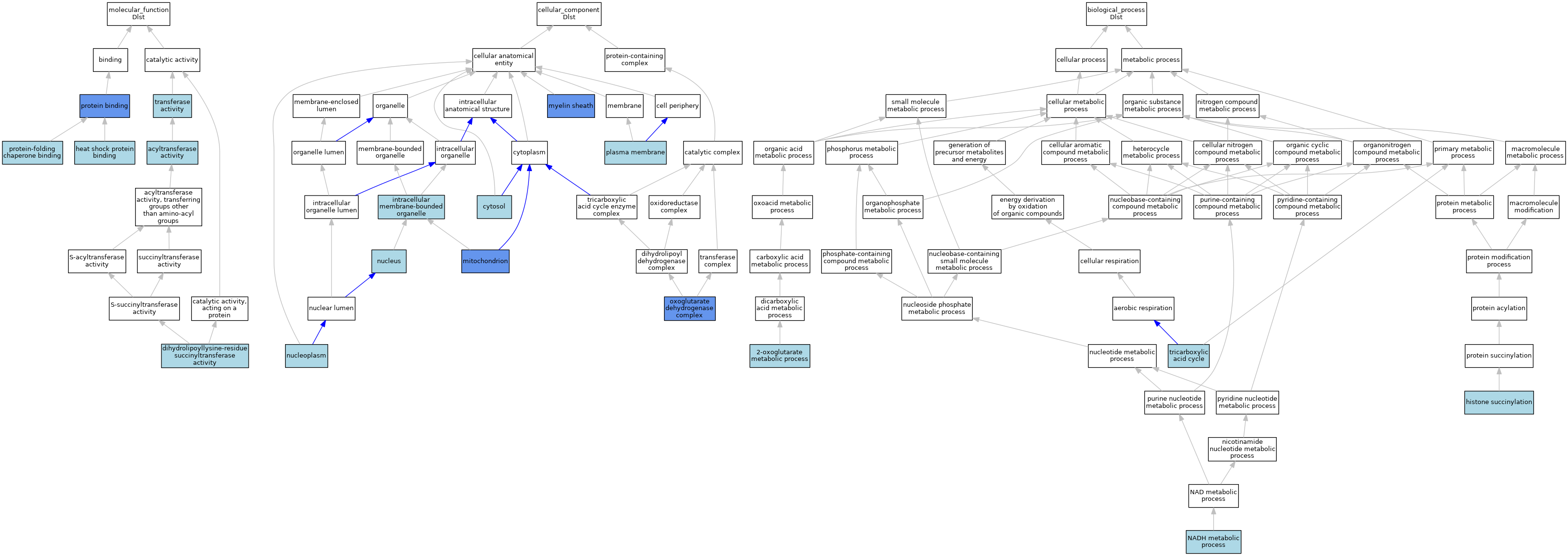
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0016746 | acyltransferase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004149 | dihydrolipoyllysine-residue succinyltransferase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004149 | dihydrolipoyllysine-residue succinyltransferase activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0004149 | dihydrolipoyllysine-residue succinyltransferase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0031072 | heat shock protein binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:263469 | |||||||||
| Molecular Function | GO:0051087 | protein-folding chaperone binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0043231 | intracellular membrane-bounded organelle | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:151002 | |||||||||
| Cellular Component | GO:0043209 | myelin sheath | HDA | J:145263 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | ISO | J:164563 | |||||||||
| Cellular Component | GO:0045252 | oxoglutarate dehydrogenase complex | ISO | J:155856 | |||||||||
| Cellular Component | GO:0045252 | oxoglutarate dehydrogenase complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0045252 | oxoglutarate dehydrogenase complex | IMP | J:188319 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:155856 | |||||||||
| Biological Process | GO:0006103 | 2-oxoglutarate metabolic process | ISO | J:155856 | |||||||||
| Biological Process | GO:0106077 | histone succinylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0006734 | NADH metabolic process | ISO | J:155856 | |||||||||
| Biological Process | GO:0006099 | tricarboxylic acid cycle | ISO | J:164563 | |||||||||
| Biological Process | GO:0006099 | tricarboxylic acid cycle | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||