|
Symbol Name ID |
Kidins220
kinase D-interacting substrate 220 MGI:1924730 |
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
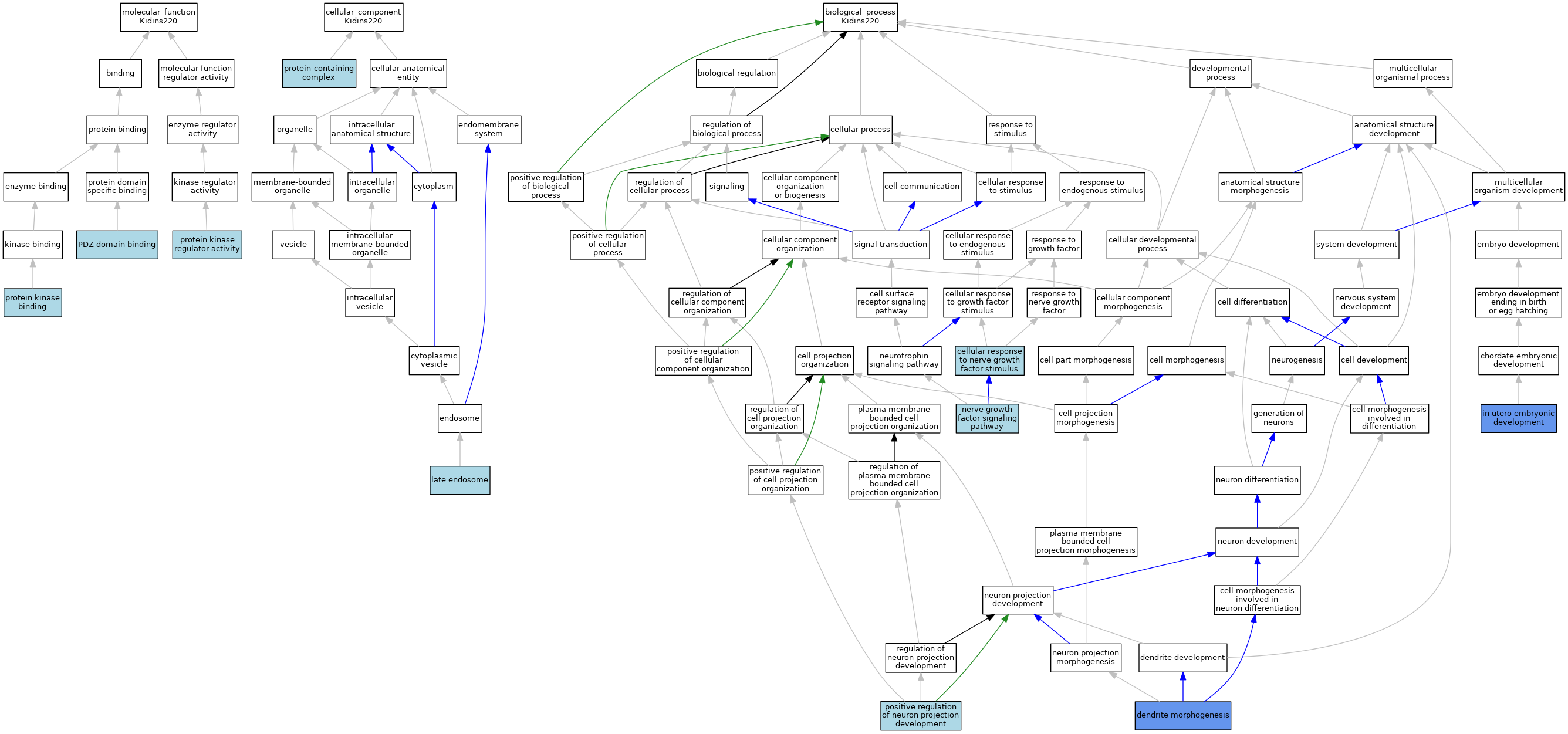
| Molecular Function | GO:0030165 | PDZ domain binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0019901 | protein kinase binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0019887 | protein kinase regulator activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0019887 | protein kinase regulator activity | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005770 | late endosome | ISO | J:155856 | |||||||||
| Cellular Component | GO:0032991 | protein-containing complex | ISO | J:155856 | |||||||||
| Biological Process | GO:1990090 | cellular response to nerve growth factor stimulus | ISO | J:155856 | |||||||||
| Biological Process | GO:0048813 | dendrite morphogenesis | IMP | J:162839 | |||||||||
| Biological Process | GO:0001701 | in utero embryonic development | IMP | J:162839 | |||||||||
| Biological Process | GO:0038180 | nerve growth factor signaling pathway | ISO | J:155856 | |||||||||
| Biological Process | GO:0038180 | nerve growth factor signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0010976 | positive regulation of neuron projection development | ISO | J:155856 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||