|
Symbol Name ID |
Elp2
elongator acetyltransferase complex subunit 2 MGI:1889642 |
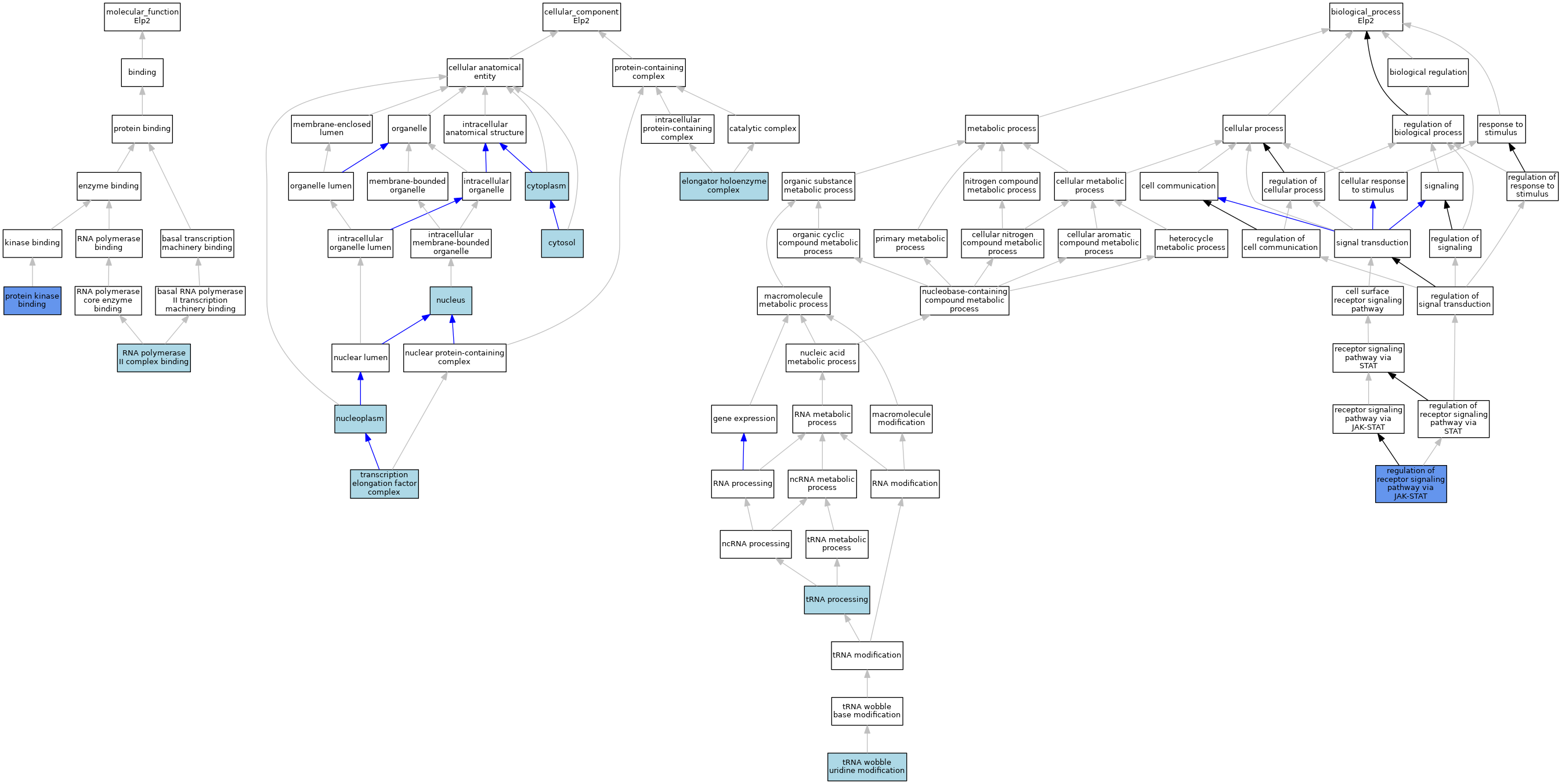
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0019901 | protein kinase binding | IDA | J:64303 | |||||||||
| Molecular Function | GO:0000993 | RNA polymerase II complex binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0033588 | elongator holoenzyme complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0033588 | elongator holoenzyme complex | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IEA | J:60000 | |||||||||
| Cellular Component | GO:0008023 | transcription elongation factor complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0008023 | transcription elongation factor complex | ISO | J:99828 | |||||||||
| Biological Process | GO:0046425 | regulation of receptor signaling pathway via JAK-STAT | IDA | J:64303 | |||||||||
| Biological Process | GO:0008033 | tRNA processing | IEA | J:60000 | |||||||||
| Biological Process | GO:0002098 | tRNA wobble uridine modification | IEA | J:72247 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||