|
Symbol Name ID |
Hnrnpdl
heterogeneous nuclear ribonucleoprotein D-like MGI:1355299 |
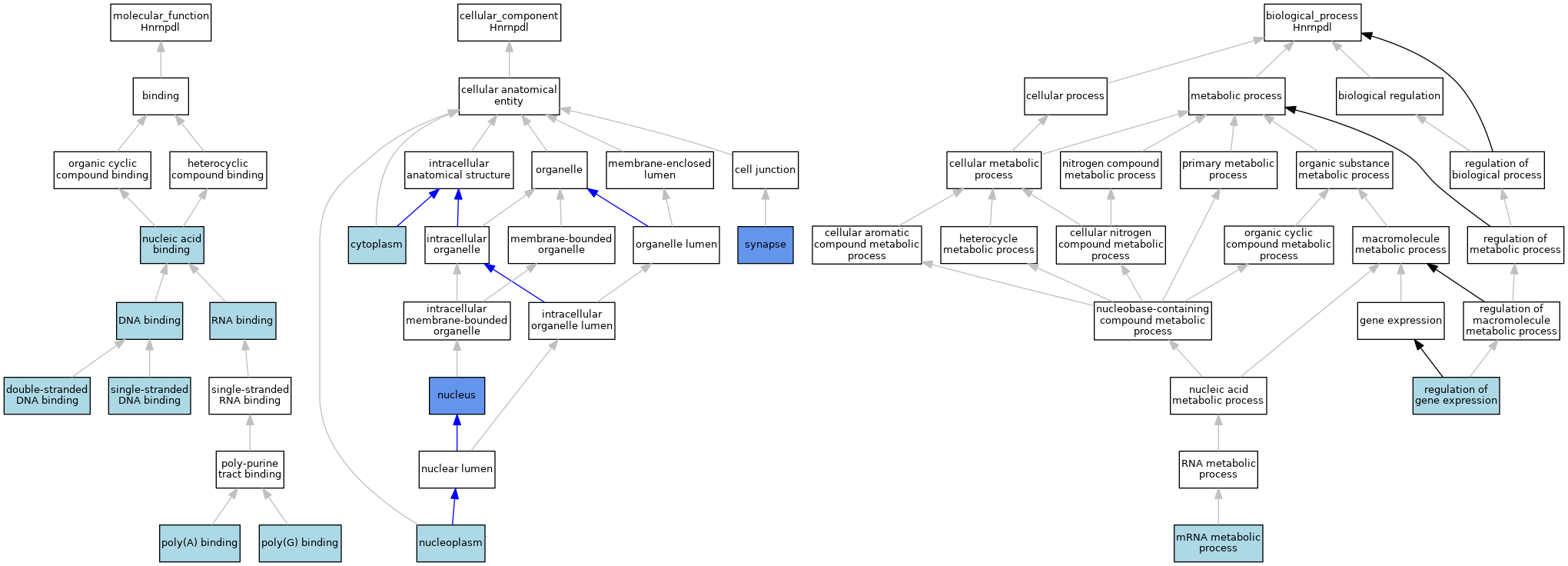
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding | ISO | J:61066 | |||||||||
| Molecular Function | GO:0003690 | double-stranded DNA binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0003676 | nucleic acid binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0008143 | poly(A) binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0008143 | poly(A) binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0034046 | poly(G) binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0034046 | poly(G) binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0003723 | RNA binding | ISO | J:61066 | |||||||||
| Molecular Function | GO:0003697 | single-stranded DNA binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005634 | nucleus | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IDA | J:61066 | |||||||||
| Cellular Component | GO:0045202 | synapse | IDA | J:263419 | |||||||||
| Cellular Component | GO:0045202 | synapse | EXP | J:263419 | |||||||||
| Biological Process | GO:0016071 | mRNA metabolic process | ISO | J:61066 | |||||||||
| Biological Process | GO:0010468 | regulation of gene expression | ISO | J:164563 | |||||||||
| Biological Process | GO:0010468 | regulation of gene expression | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||