|
Symbol Name ID |
Spo11
SPO11 initiator of meiotic double stranded breaks MGI:1349669 |
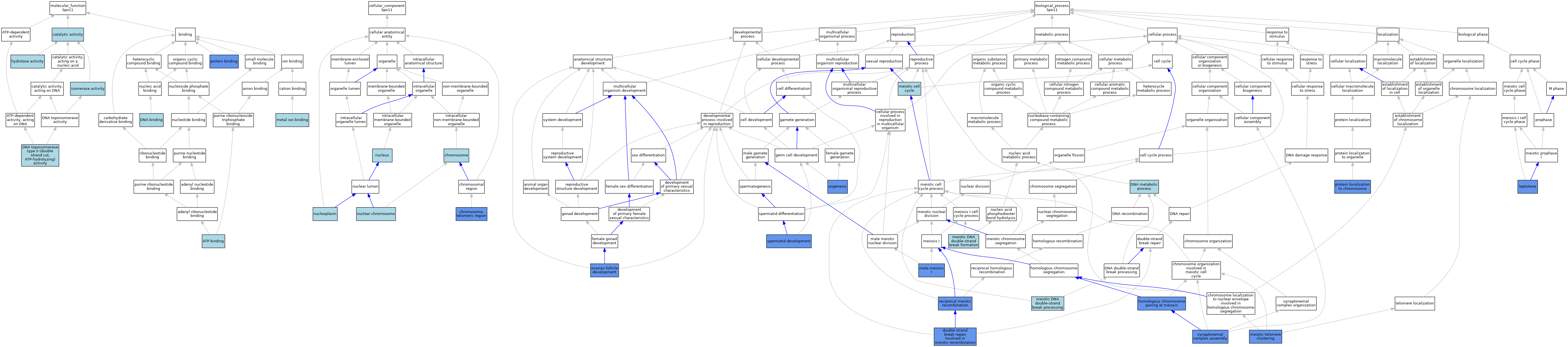
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0003824 | catalytic activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0003677 | DNA binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0003918 | DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0003918 | DNA topoisomerase type II (double strand cut, ATP-hydrolyzing) activity | IEA | J:72245 | |||||||||
| Molecular Function | GO:0016787 | hydrolase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016853 | isomerase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0046872 | metal ion binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:229524 | |||||||||
| Cellular Component | GO:0005694 | chromosome | IEA | J:72247 | |||||||||
| Cellular Component | GO:0000781 | chromosome, telomeric region | IDA | J:158887 | |||||||||
| Cellular Component | GO:0000228 | nuclear chromosome | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912344 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-MMU-912357 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IEA | J:60000 | |||||||||
| Biological Process | GO:0006259 | DNA metabolic process | IEA | J:72247 | |||||||||
| Biological Process | GO:1990918 | double-strand break repair involved in meiotic recombination | IGI | J:236226 | |||||||||
| Biological Process | GO:0007129 | homologous chromosome pairing at meiosis | IMP | J:140402 | |||||||||
| Biological Process | GO:0007129 | homologous chromosome pairing at meiosis | IMP | J:209347 | |||||||||
| Biological Process | GO:0000237 | leptotene | IMP | J:147391 | |||||||||
| Biological Process | GO:0007141 | male meiosis I | IMP | J:140402 | |||||||||
| Biological Process | GO:0007141 | male meiosis I | IMP | J:123677 | |||||||||
| Biological Process | GO:0051321 | meiotic cell cycle | IEA | J:60000 | |||||||||
| Biological Process | GO:0042138 | meiotic DNA double-strand break formation | IBA | J:265628 | |||||||||
| Biological Process | GO:0000706 | meiotic DNA double-strand break processing | IBA | J:265628 | |||||||||
| Biological Process | GO:0045141 | meiotic telomere clustering | IMP | J:147391 | |||||||||
| Biological Process | GO:0048477 | oogenesis | IMP | J:213984 | |||||||||
| Biological Process | GO:0001541 | ovarian follicle development | IMP | J:96250 | |||||||||
| Biological Process | GO:0034502 | protein localization to chromosome | IMP | J:161203 | |||||||||
| Biological Process | GO:0007131 | reciprocal meiotic recombination | IBA | J:265628 | |||||||||
| Biological Process | GO:0007131 | reciprocal meiotic recombination | IMP | J:104590 | |||||||||
| Biological Process | GO:0007286 | spermatid development | IMP | J:123677 | |||||||||
| Biological Process | GO:0007130 | synaptonemal complex assembly | IGI | J:209347 | |||||||||
| Biological Process | GO:0007130 | synaptonemal complex assembly | IMP | J:236226 | |||||||||
| Biological Process | GO:0007130 | synaptonemal complex assembly | IGI | J:209347 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||