|
Symbol Name ID |
Sra1
steroid receptor RNA activator 1 MGI:1344414 |
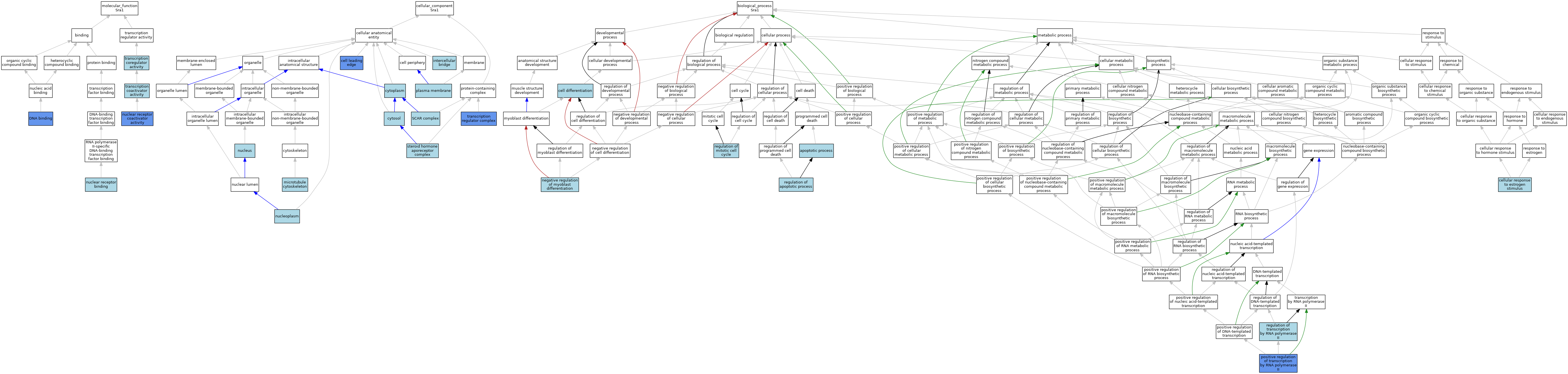
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding | IDA | J:92595 | |||||||||
| Molecular Function | GO:0016922 | nuclear receptor binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0030374 | nuclear receptor coactivator activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0030374 | nuclear receptor coactivator activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0030374 | nuclear receptor coactivator activity | IDA | J:92505 | |||||||||
| Molecular Function | GO:0003713 | transcription coactivator activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0003713 | transcription coactivator activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0003712 | transcription coregulator activity | ISO | J:54002 | |||||||||
| Cellular Component | GO:0031252 | cell leading edge | IDA | J:119028 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0045171 | intercellular bridge | ISO | J:164563 | |||||||||
| Cellular Component | GO:0015630 | microtubule cytoskeleton | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0031209 | SCAR complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005831 | steroid hormone aporeceptor complex | ISO | J:54002 | |||||||||
| Cellular Component | GO:0005667 | transcription regulator complex | IDA | J:92505 | |||||||||
| Biological Process | GO:0006915 | apoptotic process | IEA | J:60000 | |||||||||
| Biological Process | GO:0030154 | cell differentiation | ISO | J:164563 | |||||||||
| Biological Process | GO:0071391 | cellular response to estrogen stimulus | ISO | J:164563 | |||||||||
| Biological Process | GO:0045662 | negative regulation of myoblast differentiation | ISO | J:164563 | |||||||||
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | IDA | J:92505 | |||||||||
| Biological Process | GO:0042981 | regulation of apoptotic process | ISO | J:164563 | |||||||||
| Biological Process | GO:0007346 | regulation of mitotic cell cycle | ISO | J:164563 | |||||||||
| Biological Process | GO:0006357 | regulation of transcription by RNA polymerase II | ISO | J:54002 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||