|
Symbol Name ID |
Rps6ka2
ribosomal protein S6 kinase, polypeptide 2 MGI:1342290 |
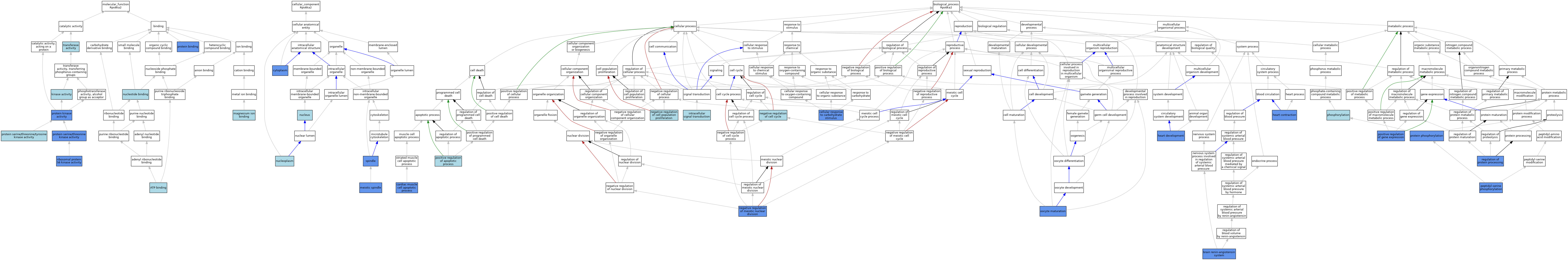
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016301 | kinase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0000287 | magnesium ion binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:94781 | |||||||||
| Molecular Function | GO:0004672 | protein kinase activity | IDA | J:94781 | |||||||||
| Molecular Function | GO:0004674 | protein serine/threonine kinase activity | TAS | Reactome:R-NUL-3876071 | |||||||||
| Molecular Function | GO:0004674 | protein serine/threonine kinase activity | IDA | J:16744 | |||||||||
| Molecular Function | GO:0004712 | protein serine/threonine/tyrosine kinase activity | ISO | J:199676 | |||||||||
| Molecular Function | GO:0004711 | ribosomal protein S6 kinase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0004711 | ribosomal protein S6 kinase activity | IDA | J:121666 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:94781 | |||||||||
| Cellular Component | GO:0072687 | meiotic spindle | IDA | J:94781 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | TAS | Reactome:R-NUL-3876071 | |||||||||
| Cellular Component | GO:0005654 | nucleoplasm | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005634 | nucleus | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005819 | spindle | IDA | J:93217 | |||||||||
| Biological Process | GO:0002035 | brain renin-angiotensin system | IDA | J:121666 | |||||||||
| Biological Process | GO:0010659 | cardiac muscle cell apoptotic process | IDA | J:121666 | |||||||||
| Biological Process | GO:0071322 | cellular response to carbohydrate stimulus | IDA | J:121666 | |||||||||
| Biological Process | GO:0060047 | heart contraction | IDA | J:121666 | |||||||||
| Biological Process | GO:0007507 | heart development | IDA | J:121666 | |||||||||
| Biological Process | GO:0035556 | intracellular signal transduction | IEA | J:72247 | |||||||||
| Biological Process | GO:0045786 | negative regulation of cell cycle | ISO | J:164563 | |||||||||
| Biological Process | GO:0008285 | negative regulation of cell population proliferation | ISO | J:164563 | |||||||||
| Biological Process | GO:0045835 | negative regulation of meiotic nuclear division | IDA | J:94781 | |||||||||
| Biological Process | GO:0001556 | oocyte maturation | IDA | J:94781 | |||||||||
| Biological Process | GO:0018105 | peptidyl-serine phosphorylation | IDA | J:16744 | |||||||||
| Biological Process | GO:0016310 | phosphorylation | IEA | J:60000 | |||||||||
| Biological Process | GO:0043065 | positive regulation of apoptotic process | ISO | J:164563 | |||||||||
| Biological Process | GO:0010628 | positive regulation of gene expression | IDA | J:121666 | |||||||||
| Biological Process | GO:0006468 | protein phosphorylation | IDA | J:94781 | |||||||||
| Biological Process | GO:0070613 | regulation of protein processing | IMP | J:121666 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||