|
Source |
Alliance of Genome Resources |
| Category | ID | Classification term | Gene | Evidence | Inferred from | Organism | Reference |
|---|---|---|---|---|---|---|---|
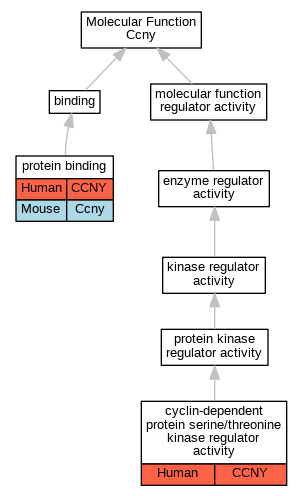
| Molecular Function | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity | CCNY | IDA | Human | PMID:19524571 | |
| Molecular Function | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity | CCNY | IDA | Human | PMID:20059949 | |
| Molecular Function | GO:0005515 | protein binding | CCNY | IPI | UniProtKB:O94921 | Human | PMID:19524571 |
| Molecular Function | GO:0005515 | protein binding | CCNY | IPI | UniProtKB:O75581 | Human | PMID:20059949 |
| Molecular Function | GO:0005515 | protein binding | CCNY | IPI | UniProtKB:O94921 | Human | PMID:20059949 |
| Molecular Function | GO:0005515 | protein binding | CCNY | IPI | UniProtKB:Q13616 | Human | PMID:24794231 |
| Molecular Function | GO:0005515 | protein binding | Ccny | IPI | UniProtKB:Q04735 | Mouse | MGI:MGI:5319028|PMID:22184064 |
| Molecular Function | GO:0005515 | protein binding | Ccny | IPI | UniProtKB:Q04735 | Mouse | MGI:MGI:5688365|PMID:26305884 |
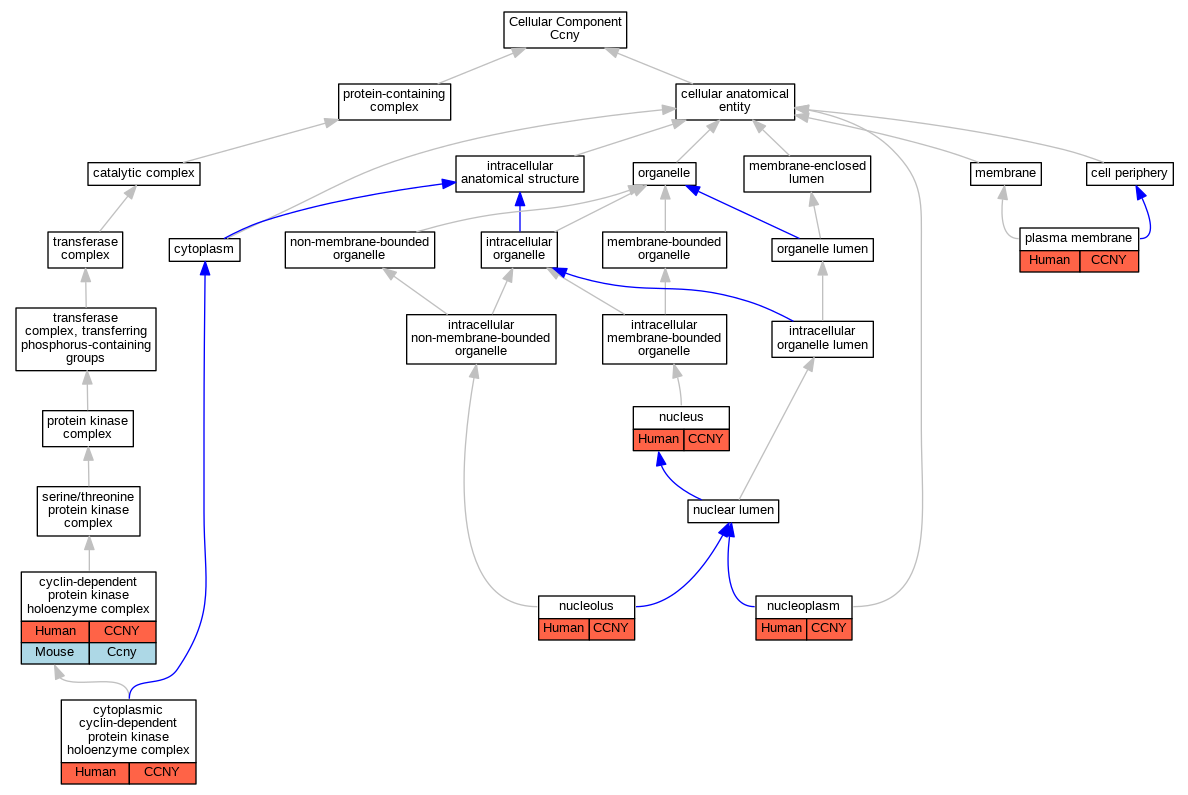
| Cellular Component | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex | CCNY | IPI | Human | PMID:19524571 | |
| Cellular Component | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex | Ccny | IPI | Mouse | MGI:MGI:3851889|PMID:19524571 | |
| Cellular Component | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex | CCNY | IDA | Human | PMID:19524571 | |
| Cellular Component | GO:0000308 | cytoplasmic cyclin-dependent protein kinase holoenzyme complex | CCNY | IDA | Human | PMID:20059949 | |
| Cellular Component | GO:0005730 | nucleolus | CCNY | IDA | Human | GO_REF:0000052 | |
| Cellular Component | GO:0005654 | nucleoplasm | CCNY | IDA | Human | GO_REF:0000052 | |
| Cellular Component | GO:0005634 | nucleus | CCNY | IDA | Human | GO_REF:0000054 | |
| Cellular Component | GO:0005886 | plasma membrane | CCNY | IDA | Human | PMID:19524571 | |
| Cellular Component | GO:0005886 | plasma membrane | CCNY | IDA | Human | PMID:20059949 | |
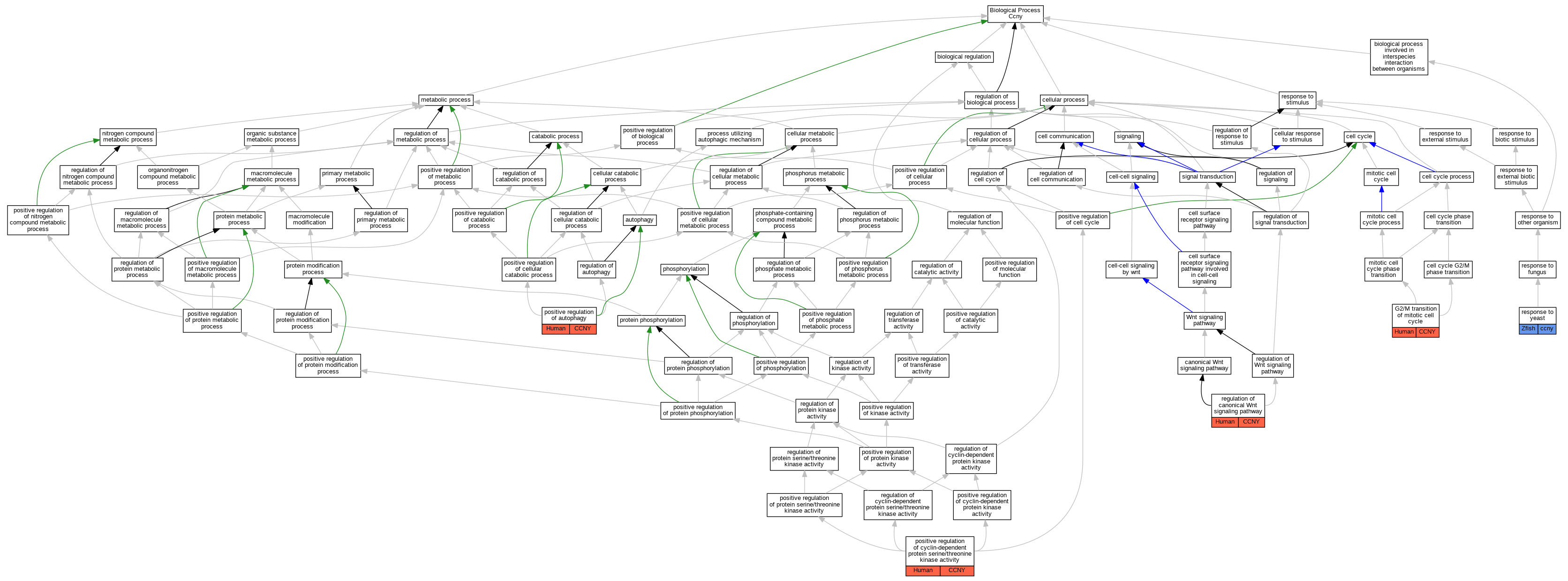
| Biological Process | GO:0000086 | G2/M transition of mitotic cell cycle | CCNY | IDA | Human | PMID:20059949 | |
| Biological Process | GO:0010508 | positive regulation of autophagy | CCNY | IDA | Human | PMID:32098961 | |
| Biological Process | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity | CCNY | IDA | Human | PMID:19524571 | |
| Biological Process | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity | CCNY | IDA | Human | PMID:20059949 | |
| Biological Process | GO:0060828 | regulation of canonical Wnt signaling pathway | CCNY | IDA | Human | PMID:20059949 | |
| Biological Process | GO:0001878 | response to yeast | ccny | IDA | Zfish | PMID:23947337 |
| EXP Inferred from experiment |
| IDA Inferred from direct assay |
| IEP Inferred from expression pattern |
| IGI Inferred from genetic interaction |
| IMP Inferred from mutant phenotype |
| IPI Inferred from physical interaction |
| HTP Inferred from High Throughput Experiment |
| HDA Inferred from High Throughput Direct Assay |
| HMP Inferred from High Throughput Mutant Phenotype |
| HGI Inferred from High Throughput Genetic Interaction |
| HEP Inferred from High Throughput Expression Pattern |
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||