|
Symbol Name ID |
Oprl1
opioid receptor-like 1 MGI:97440 |
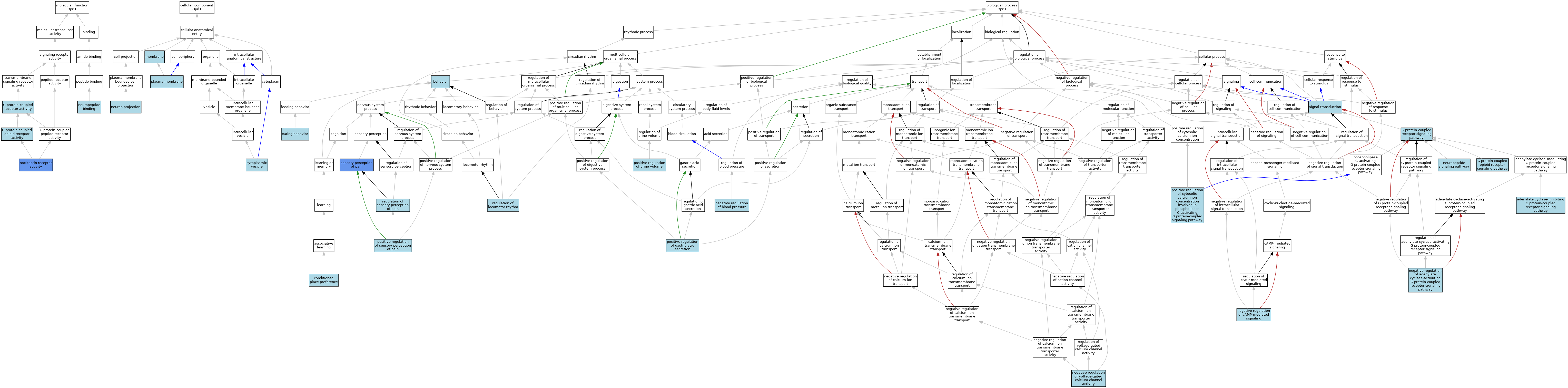
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0004985 | G protein-coupled opioid receptor activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0004930 | G protein-coupled receptor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0042923 | neuropeptide binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0001626 | nociceptin receptor activity | IMP | J:102658 | |||||||||
| Molecular Function | GO:0001626 | nociceptin receptor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0001626 | nociceptin receptor activity | IBA | J:265628 | |||||||||
| Cellular Component | GO:0031410 | cytoplasmic vesicle | IEA | J:60000 | |||||||||
| Cellular Component | GO:0016020 | membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0043005 | neuron projection | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Biological Process | GO:0007193 | adenylate cyclase-inhibiting G protein-coupled receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0007610 | behavior | IEA | J:60000 | |||||||||
| Biological Process | GO:1990708 | conditioned place preference | ISO | J:155856 | |||||||||
| Biological Process | GO:0042755 | eating behavior | ISO | J:155856 | |||||||||
| Biological Process | GO:0038003 | G protein-coupled opioid receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0007186 | G protein-coupled receptor signaling pathway | IEA | J:60000 | |||||||||
| Biological Process | GO:0007186 | G protein-coupled receptor signaling pathway | IEA | J:72247 | |||||||||
| Biological Process | GO:0106072 | negative regulation of adenylate cyclase-activating G protein-coupled receptor signaling pathway | ISO | J:155856 | |||||||||
| Biological Process | GO:0045776 | negative regulation of blood pressure | ISO | J:155856 | |||||||||
| Biological Process | GO:0043951 | negative regulation of cAMP-mediated signaling | ISO | J:155856 | |||||||||
| Biological Process | GO:1901386 | negative regulation of voltage-gated calcium channel activity | ISO | J:155856 | |||||||||
| Biological Process | GO:0007218 | neuropeptide signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G protein-coupled signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0060454 | positive regulation of gastric acid secretion | ISO | J:155856 | |||||||||
| Biological Process | GO:1904058 | positive regulation of sensory perception of pain | ISO | J:155856 | |||||||||
| Biological Process | GO:0035810 | positive regulation of urine volume | ISO | J:155856 | |||||||||
| Biological Process | GO:1904059 | regulation of locomotor rhythm | ISO | J:155856 | |||||||||
| Biological Process | GO:0051930 | regulation of sensory perception of pain | ISO | J:155856 | |||||||||
| Biological Process | GO:0019233 | sensory perception of pain | IMP | J:102658 | |||||||||
| Biological Process | GO:0019233 | sensory perception of pain | IBA | J:265628 | |||||||||
| Biological Process | GO:0007165 | signal transduction | IEA | J:60000 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||