|
Symbol Name ID |
Glud1
glutamate dehydrogenase 1 MGI:95753 |
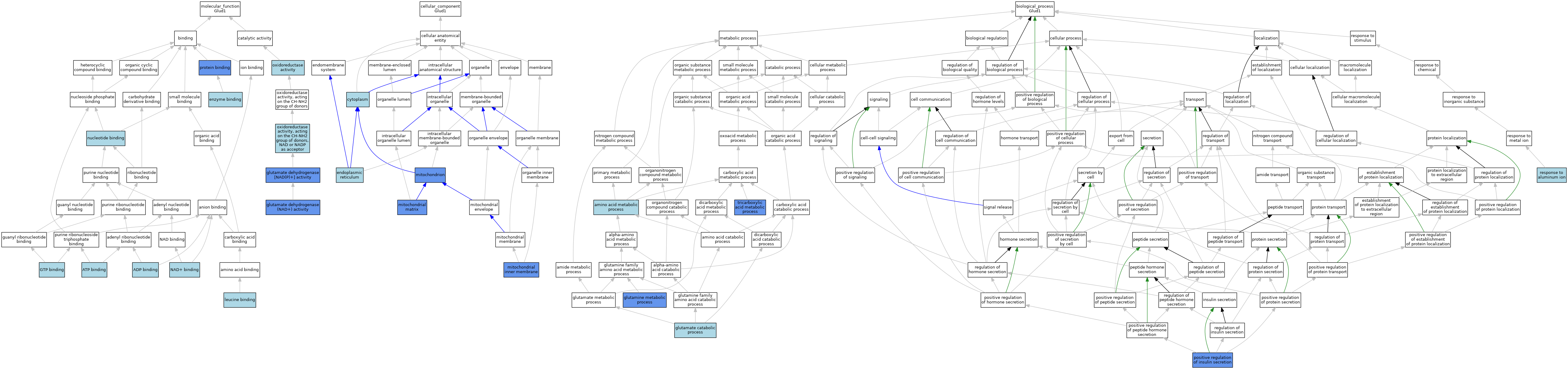
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0043531 | ADP binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0043531 | ADP binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0019899 | enzyme binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0004352 | glutamate dehydrogenase (NAD+) activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004352 | glutamate dehydrogenase (NAD+) activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004352 | glutamate dehydrogenase (NAD+) activity | IDA | J:166803 | |||||||||
| Molecular Function | GO:0004352 | glutamate dehydrogenase (NAD+) activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0004352 | glutamate dehydrogenase (NAD+) activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0004353 | glutamate dehydrogenase [NAD(P)+] activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004353 | glutamate dehydrogenase [NAD(P)+] activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004353 | glutamate dehydrogenase [NAD(P)+] activity | IDA | J:198927 | |||||||||
| Molecular Function | GO:0004353 | glutamate dehydrogenase [NAD(P)+] activity | IDA | J:129427 | |||||||||
| Molecular Function | GO:0004353 | glutamate dehydrogenase [NAD(P)+] activity | IDA | J:233904 | |||||||||
| Molecular Function | GO:0004353 | glutamate dehydrogenase [NAD(P)+] activity | IDA | J:73910 | |||||||||
| Molecular Function | GO:0005525 | GTP binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0005525 | GTP binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0070728 | leucine binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0070728 | leucine binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0070403 | NAD+ binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016491 | oxidoreductase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016491 | oxidoreductase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016639 | oxidoreductase activity, acting on the CH-NH2 group of donors, NAD or NADP as acceptor | IEA | J:72247 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:166803 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:129427 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:200967 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005783 | endoplasmic reticulum | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005743 | mitochondrial inner membrane | HDA | J:100953 | |||||||||
| Cellular Component | GO:0005759 | mitochondrial matrix | IDA | J:129009 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IDA | J:233904 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:151002 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IDA | J:129427 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Biological Process | GO:0006520 | amino acid metabolic process | IEA | J:72247 | |||||||||
| Biological Process | GO:0006538 | glutamate catabolic process | IBA | J:265628 | |||||||||
| Biological Process | GO:0006541 | glutamine metabolic process | IMP | J:198927 | |||||||||
| Biological Process | GO:0032024 | positive regulation of insulin secretion | IGI | J:129427 | |||||||||
| Biological Process | GO:0010044 | response to aluminum ion | ISO | J:155856 | |||||||||
| Biological Process | GO:0072350 | tricarboxylic acid metabolic process | IMP | J:198927 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||