|
Symbol Name ID |
Drd5
dopamine receptor D5 MGI:94927 |
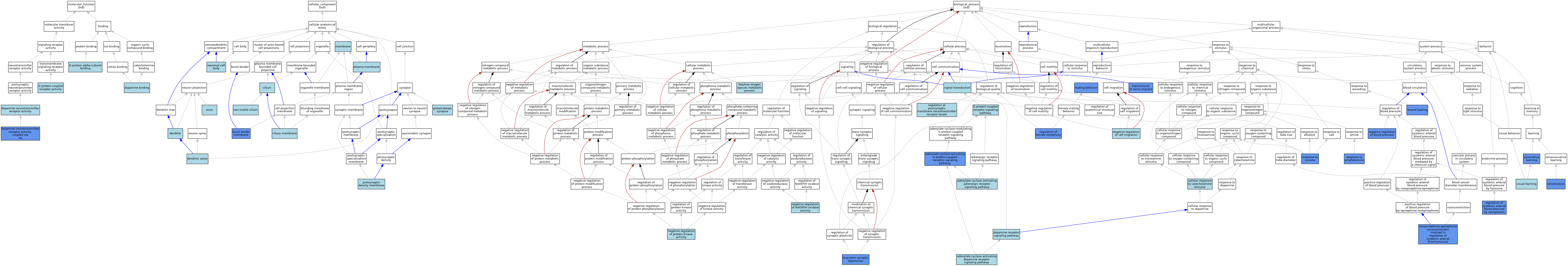
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0035240 | dopamine binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004952 | dopamine neurotransmitter receptor activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0004952 | dopamine neurotransmitter receptor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004952 | dopamine neurotransmitter receptor activity | ISO | J:73065 | |||||||||
| Molecular Function | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs | IBA | J:265628 | |||||||||
| Molecular Function | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs | IDA | J:114543 | |||||||||
| Molecular Function | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs | IDA | J:114543 | |||||||||
| Molecular Function | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs | IMP | J:100026 | |||||||||
| Molecular Function | GO:0004930 | G protein-coupled receptor activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0001965 | G-protein alpha-subunit binding | ISO | J:155856 | |||||||||
| Cellular Component | GO:0030424 | axon | ISO | J:155856 | |||||||||
| Cellular Component | GO:0031526 | brush border membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0031526 | brush border membrane | IDA | J:95570 | |||||||||
| Cellular Component | GO:0060170 | ciliary membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005929 | cilium | ISO | J:246963 | |||||||||
| Cellular Component | GO:0030425 | dendrite | ISO | J:155856 | |||||||||
| Cellular Component | GO:0043197 | dendritic spine | ISO | J:155856 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:72247 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0043025 | neuronal cell body | ISO | J:155856 | |||||||||
| Cellular Component | GO:0097730 | non-motile cilium | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0098839 | postsynaptic density membrane | ISO | J:155856 | |||||||||
| Biological Process | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway | ISO | J:155856 | |||||||||
| Biological Process | GO:0007189 | adenylate cyclase-activating G protein-coupled receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0007189 | adenylate cyclase-activating G protein-coupled receptor signaling pathway | ISO | J:73065 | |||||||||
| Biological Process | GO:0007189 | adenylate cyclase-activating G protein-coupled receptor signaling pathway | IDA | J:95570 | |||||||||
| Biological Process | GO:0008306 | associative learning | IMP | J:114239 | |||||||||
| Biological Process | GO:0071870 | cellular response to catecholamine stimulus | ISO | J:164563 | |||||||||
| Biological Process | GO:0007212 | dopamine receptor signaling pathway | ISO | J:155856 | |||||||||
| Biological Process | GO:0007212 | dopamine receptor signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0007186 | G protein-coupled receptor signaling pathway | IEA | J:60000 | |||||||||
| Biological Process | GO:0007186 | G protein-coupled receptor signaling pathway | IEA | J:72247 | |||||||||
| Biological Process | GO:0060292 | long-term synaptic depression | IMP | J:100026 | |||||||||
| Biological Process | GO:0007617 | mating behavior | IMP | J:114239 | |||||||||
| Biological Process | GO:0045776 | negative regulation of blood pressure | IGI | J:95908 | |||||||||
| Biological Process | GO:0045776 | negative regulation of blood pressure | IMP | J:95570 | |||||||||
| Biological Process | GO:0045776 | negative regulation of blood pressure | IMP | J:95908 | |||||||||
| Biological Process | GO:0030336 | negative regulation of cell migration | ISO | J:155856 | |||||||||
| Biological Process | GO:0033861 | negative regulation of NAD(P)H oxidase activity | ISO | J:164563 | |||||||||
| Biological Process | GO:0006469 | negative regulation of protein kinase activity | ISO | J:155856 | |||||||||
| Biological Process | GO:0001994 | norepinephrine-epinephrine vasoconstriction involved in regulation of systemic arterial blood pressure | IMP | J:95908 | |||||||||
| Biological Process | GO:0072593 | reactive oxygen species metabolic process | ISO | J:164563 | |||||||||
| Biological Process | GO:0045924 | regulation of female receptivity | IMP | J:114239 | |||||||||
| Biological Process | GO:0099072 | regulation of postsynaptic membrane neurotransmitter receptor levels | ISO | J:155856 | |||||||||
| Biological Process | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin | IGI | J:95908 | |||||||||
| Biological Process | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin | IGI | J:95908 | |||||||||
| Biological Process | GO:0001975 | response to amphetamine | IGI | J:103950 | |||||||||
| Biological Process | GO:0042220 | response to cocaine | IMP | J:103887 | |||||||||
| Biological Process | GO:0046960 | sensitization | IGI | J:103950 | |||||||||
| Biological Process | GO:0007165 | signal transduction | IEA | J:60000 | |||||||||
| Biological Process | GO:0019226 | transmission of nerve impulse | IMP | J:84475 | |||||||||
| Biological Process | GO:0008542 | visual learning | ISO | J:155856 | |||||||||
| Biological Process | GO:0042060 | wound healing | IMP | J:111218 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||