|
Symbol Name ID |
Kyat3
kynurenine aminotransferase 3 MGI:2677849 |
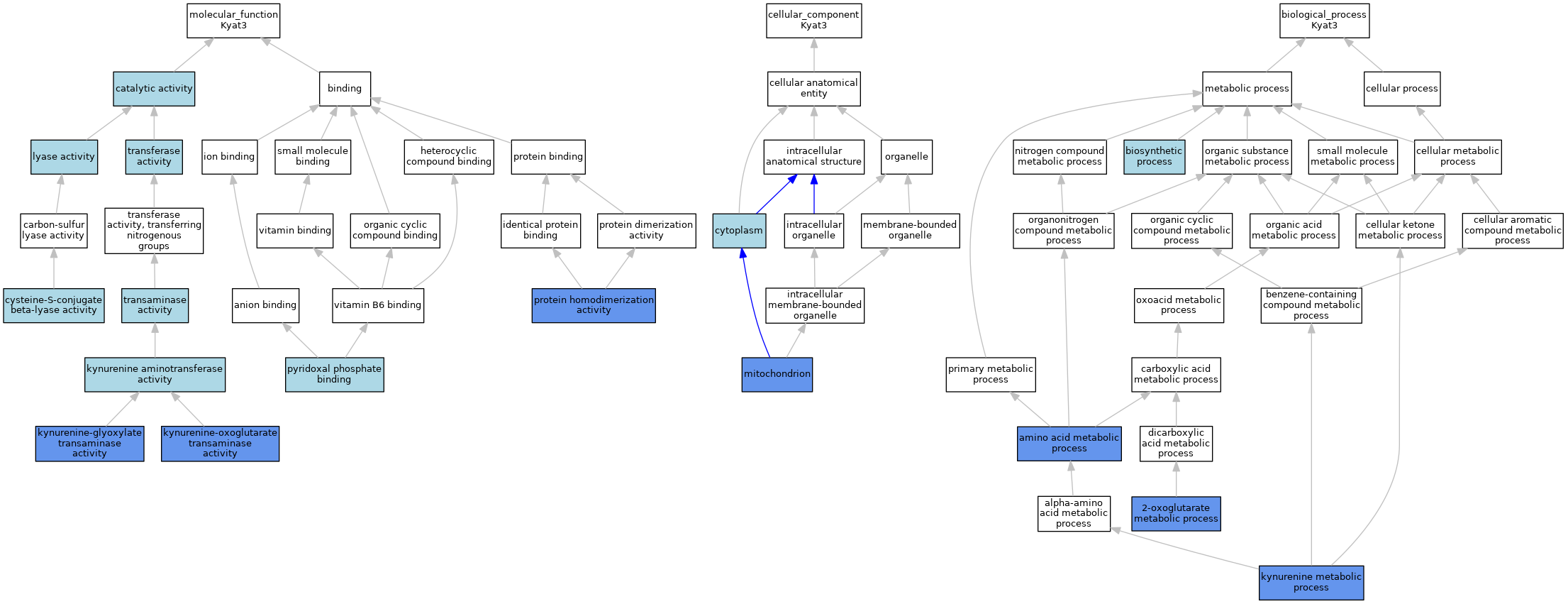
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0047804 | cysteine-S-conjugate beta-lyase activity | IEA | J:72245 | |||||||||
| Molecular Function | GO:0036137 | kynurenine aminotransferase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0047315 | kynurenine-glyoxylate transaminase activity | IDA | J:145553 | |||||||||
| Molecular Function | GO:0016212 | kynurenine-oxoglutarate transaminase activity | IDA | J:145553 | |||||||||
| Molecular Function | GO:0016212 | kynurenine-oxoglutarate transaminase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0016829 | lyase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0042803 | protein homodimerization activity | IPI | J:145553 | |||||||||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0008483 | transaminase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:151002 | |||||||||
| Biological Process | GO:0006103 | 2-oxoglutarate metabolic process | IDA | J:145553 | |||||||||
| Biological Process | GO:0006520 | amino acid metabolic process | IDA | J:145553 | |||||||||
| Biological Process | GO:0009058 | biosynthetic process | IEA | J:72247 | |||||||||
| Biological Process | GO:0070189 | kynurenine metabolic process | IDA | J:145553 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||