|
Symbol Name ID |
N4bp1
NEDD4 binding protein 1 MGI:2136825 |
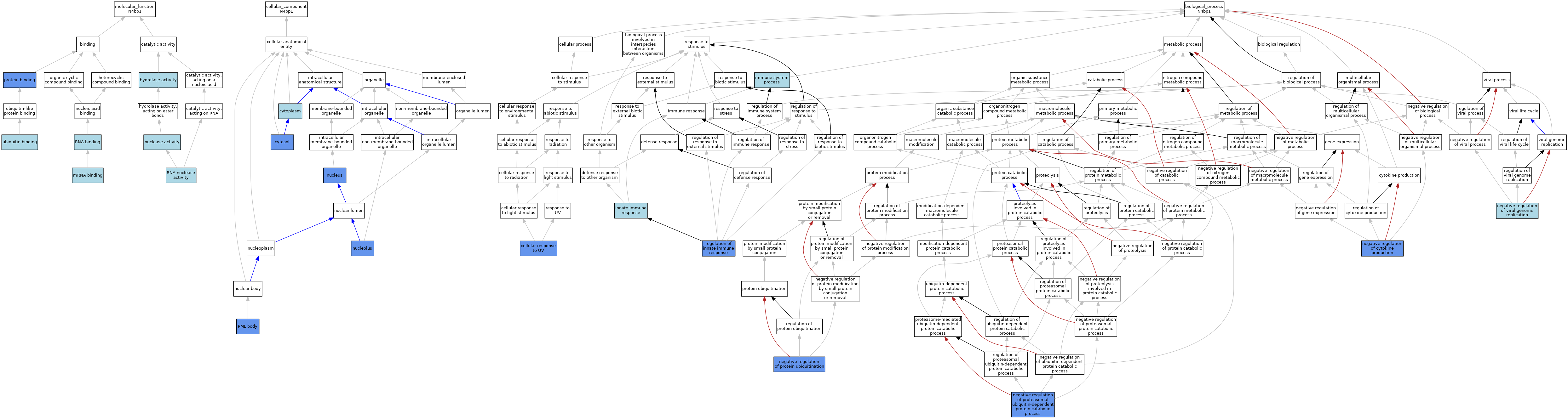
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0016787 | hydrolase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0003729 | mRNA binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004518 | nuclease activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:143823 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:164713 | |||||||||
| Molecular Function | GO:0003723 | RNA binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0003723 | RNA binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0004540 | RNA nuclease activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0043130 | ubiquitin binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005829 | cytosol | IDA | J:297419 | |||||||||
| Cellular Component | GO:0005730 | nucleolus | IDA | J:159635 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IDA | J:164713 | |||||||||
| Cellular Component | GO:0016605 | PML body | IDA | J:159635 | |||||||||
| Cellular Component | GO:0016605 | PML body | IBA | J:265628 | |||||||||
| Biological Process | GO:0034644 | cellular response to UV | IMP | J:143823 | |||||||||
| Biological Process | GO:0034644 | cellular response to UV | ISO | J:164563 | |||||||||
| Biological Process | GO:0002376 | immune system process | IEA | J:60000 | |||||||||
| Biological Process | GO:0045087 | innate immune response | IEA | J:60000 | |||||||||
| Biological Process | GO:0001818 | negative regulation of cytokine production | IMP | J:297419 | |||||||||
| Biological Process | GO:0032435 | negative regulation of proteasomal ubiquitin-dependent protein catabolic process | IDA | J:143823 | |||||||||
| Biological Process | GO:0032435 | negative regulation of proteasomal ubiquitin-dependent protein catabolic process | IBA | J:265628 | |||||||||
| Biological Process | GO:0031397 | negative regulation of protein ubiquitination | IDA | J:143823 | |||||||||
| Biological Process | GO:0031397 | negative regulation of protein ubiquitination | IBA | J:265628 | |||||||||
| Biological Process | GO:0045071 | negative regulation of viral genome replication | ISO | J:164563 | |||||||||
| Biological Process | GO:0045088 | regulation of innate immune response | IDA | J:297419 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 05/07/2024 MGI 6.23 |

|
|
|
||