|
Symbol Name ID |
Yars2
tyrosyl-tRNA synthetase 2 (mitochondrial) MGI:1917370 |
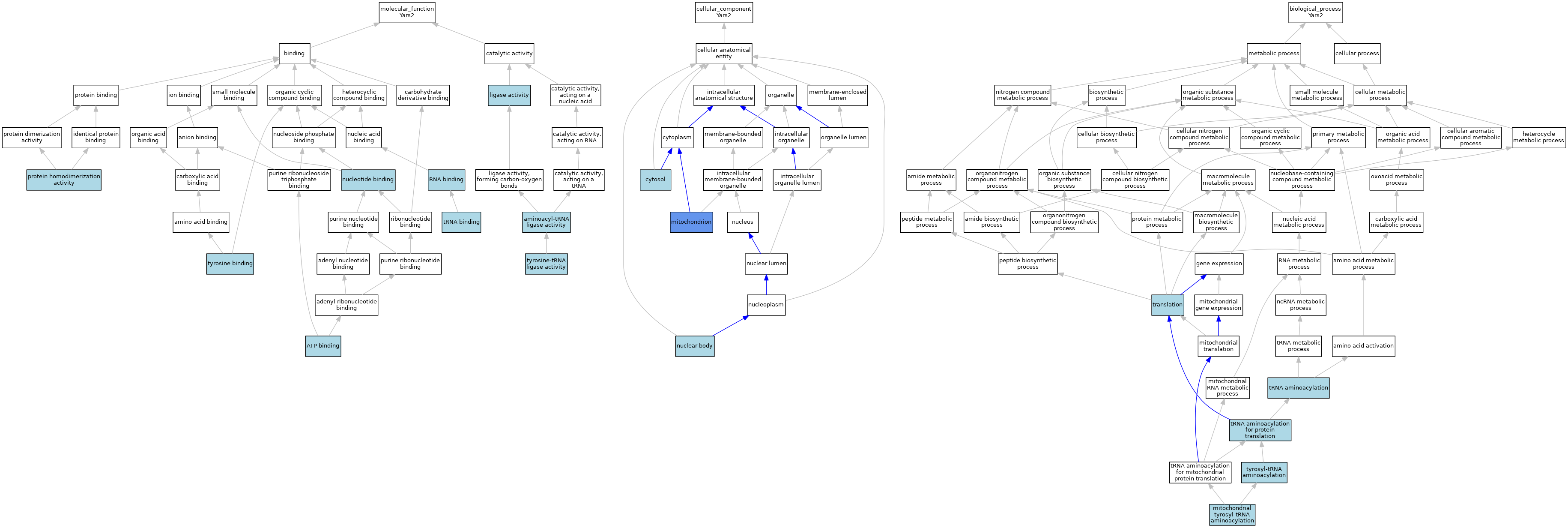
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004812 | aminoacyl-tRNA ligase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016874 | ligase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0042803 | protein homodimerization activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0003723 | RNA binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0000049 | tRNA binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0072545 | tyrosine binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004831 | tyrosine-tRNA ligase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004831 | tyrosine-tRNA ligase activity | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005829 | cytosol | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:151002 | |||||||||
| Cellular Component | GO:0016604 | nuclear body | ISO | J:164563 | |||||||||
| Biological Process | GO:0070184 | mitochondrial tyrosyl-tRNA aminoacylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0006412 | translation | IEA | J:60000 | |||||||||
| Biological Process | GO:0043039 | tRNA aminoacylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0043039 | tRNA aminoacylation | IBA | J:265628 | |||||||||
| Biological Process | GO:0006418 | tRNA aminoacylation for protein translation | IEA | J:72247 | |||||||||
| Biological Process | GO:0006437 | tyrosyl-tRNA aminoacylation | IEA | J:72247 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||