|
Symbol Name ID |
Grm4
glutamate receptor, metabotropic 4 MGI:1351341 |
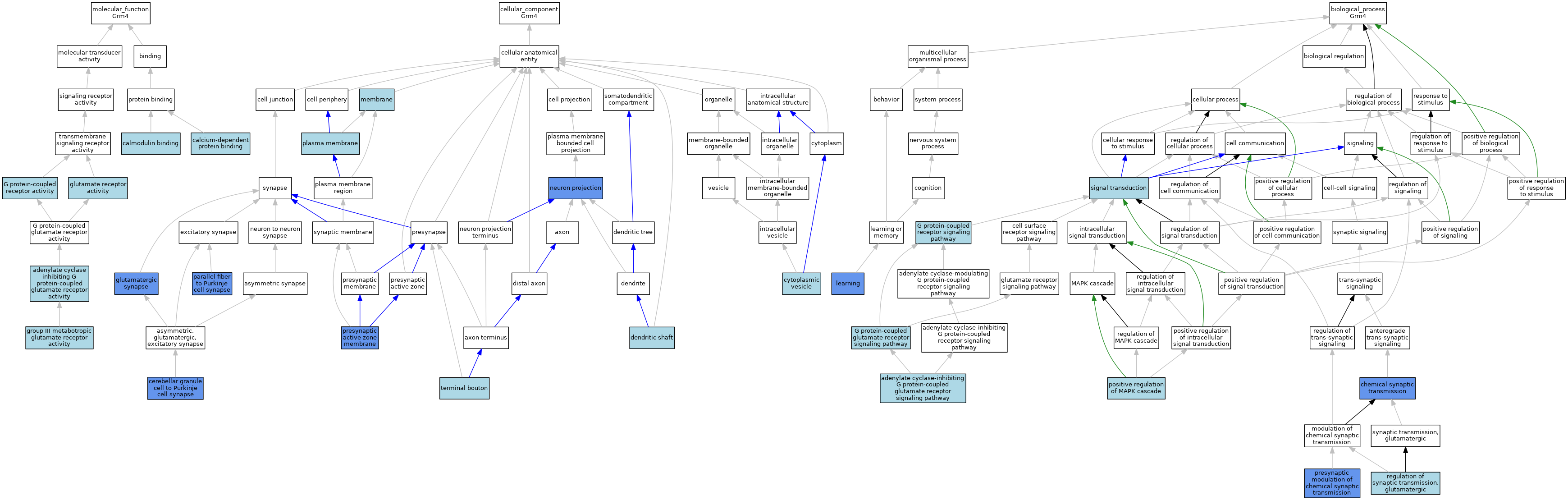
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0001640 | adenylate cyclase inhibiting G protein-coupled glutamate receptor activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0048306 | calcium-dependent protein binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0005516 | calmodulin binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0004930 | G protein-coupled receptor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0008066 | glutamate receptor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0001642 | group III metabotropic glutamate receptor activity | TAS | J:35793 | |||||||||
| Cellular Component | GO:0150048 | cerebellar granule cell to Purkinje cell synapse | IDA | J:326657 | |||||||||
| Cellular Component | GO:0150048 | cerebellar granule cell to Purkinje cell synapse | IDA | J:326657 | |||||||||
| Cellular Component | GO:0150048 | cerebellar granule cell to Purkinje cell synapse | IDA | J:326657 | |||||||||
| Cellular Component | GO:0031410 | cytoplasmic vesicle | ISO | J:164563 | |||||||||
| Cellular Component | GO:0043198 | dendritic shaft | ISO | J:155856 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:263466 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:263466 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:263466 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:263466 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:263466 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:263466 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:35793 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | EXP | J:35793 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:35793 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IMP | J:35793 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:326657 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:326657 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:326657 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:72247 | |||||||||
| Cellular Component | GO:0043005 | neuron projection | IDA | J:184110 | |||||||||
| Cellular Component | GO:0098688 | parallel fiber to Purkinje cell synapse | IDA | J:35793 | |||||||||
| Cellular Component | GO:0098688 | parallel fiber to Purkinje cell synapse | EXP | J:35793 | |||||||||
| Cellular Component | GO:0098688 | parallel fiber to Purkinje cell synapse | IDA | J:35793 | |||||||||
| Cellular Component | GO:0098688 | parallel fiber to Purkinje cell synapse | IMP | J:35793 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:263466 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:263466 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:263466 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:263466 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:263466 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:263466 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:326657 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:326657 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:326657 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:326657 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:326657 | |||||||||
| Cellular Component | GO:0048787 | presynaptic active zone membrane | IDA | J:326657 | |||||||||
| Cellular Component | GO:0043195 | terminal bouton | ISO | J:155856 | |||||||||
| Biological Process | GO:0007196 | adenylate cyclase-inhibiting G protein-coupled glutamate receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0007268 | chemical synaptic transmission | IMP | J:35793 | |||||||||
| Biological Process | GO:0007216 | G protein-coupled glutamate receptor signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0007186 | G protein-coupled receptor signaling pathway | IEA | J:60000 | |||||||||
| Biological Process | GO:0007186 | G protein-coupled receptor signaling pathway | IEA | J:72247 | |||||||||
| Biological Process | GO:0007612 | learning | IMP | J:35793 | |||||||||
| Biological Process | GO:0043410 | positive regulation of MAPK cascade | ISO | J:164563 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | IDA | J:35793 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | IDA | J:35793 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | IDA | J:35793 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | IMP | J:35793 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | IMP | J:35793 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | IDA | J:35793 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | EXP | J:35793 | |||||||||
| Biological Process | GO:0099171 | presynaptic modulation of chemical synaptic transmission | EXP | J:35793 | |||||||||
| Biological Process | GO:0051966 | regulation of synaptic transmission, glutamatergic | IBA | J:265628 | |||||||||
| Biological Process | GO:0007165 | signal transduction | IEA | J:60000 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||