|
Symbol Name ID |
Dpagt1
dolichyl-phosphate N-acetylglucosaminephosphotransferase 1 MGI:1196396 |
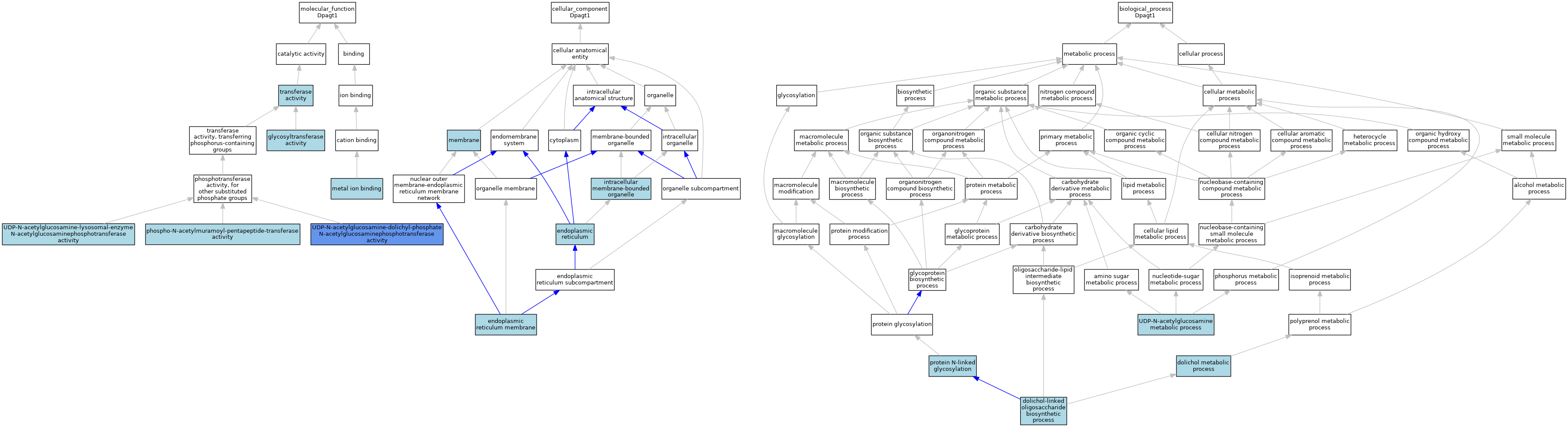
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0016757 | glycosyltransferase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0046872 | metal ion binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0008963 | phospho-N-acetylmuramoyl-pentapeptide-transferase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0003975 | UDP-N-acetylglucosamine-dolichyl-phosphate N-acetylglucosaminephosphotransferase activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0003975 | UDP-N-acetylglucosamine-dolichyl-phosphate N-acetylglucosaminephosphotransferase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0003975 | UDP-N-acetylglucosamine-dolichyl-phosphate N-acetylglucosaminephosphotransferase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0003975 | UDP-N-acetylglucosamine-dolichyl-phosphate N-acetylglucosaminephosphotransferase activity | IDA | J:1728 | |||||||||
| Molecular Function | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity | ISO | J:47505 | |||||||||
| Molecular Function | GO:0003976 | UDP-N-acetylglucosamine-lysosomal-enzyme N-acetylglucosaminephosphotransferase activity | ISS | J:1728 | |||||||||
| Cellular Component | GO:0005783 | endoplasmic reticulum | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005789 | endoplasmic reticulum membrane | TAS | J:1728 | |||||||||
| Cellular Component | GO:0043231 | intracellular membrane-bounded organelle | ISO | J:164563 | |||||||||
| Cellular Component | GO:0043231 | intracellular membrane-bounded organelle | ISO | J:155856 | |||||||||
| Cellular Component | GO:0016020 | membrane | ISO | J:164563 | |||||||||
| Biological Process | GO:0019348 | dolichol metabolic process | ISO | J:155856 | |||||||||
| Biological Process | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process | ISO | J:155856 | |||||||||
| Biological Process | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process | ISO | J:164563 | |||||||||
| Biological Process | GO:0006488 | dolichol-linked oligosaccharide biosynthetic process | TAS | J:59695 | |||||||||
| Biological Process | GO:0006487 | protein N-linked glycosylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0006047 | UDP-N-acetylglucosamine metabolic process | ISO | J:155856 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||