|
Symbol Name ID |
Sftpd
surfactant associated protein D MGI:109515 |
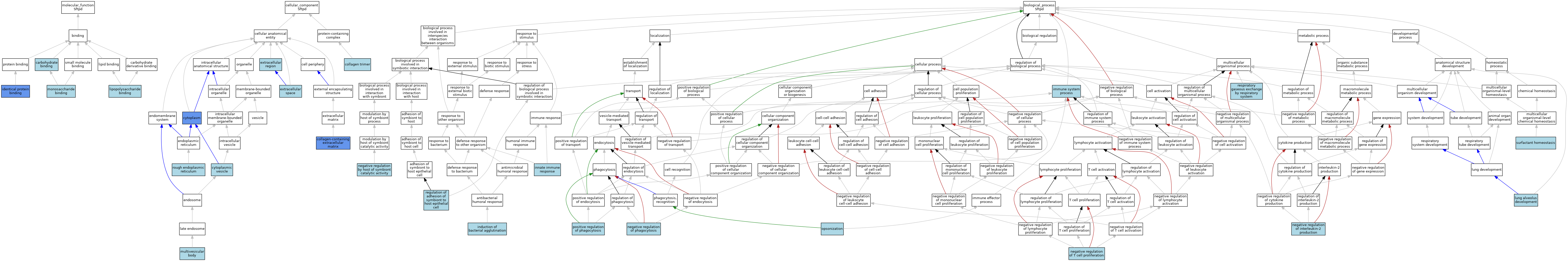
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0030246 | carbohydrate binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0042802 | identical protein binding | IPI | J:144759 | |||||||||
| Molecular Function | GO:0042802 | identical protein binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0001530 | lipopolysaccharide binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0048029 | monosaccharide binding | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005581 | collagen trimer | IEA | J:60000 | |||||||||
| Cellular Component | GO:0062023 | collagen-containing extracellular matrix | HDA | J:265604 | |||||||||
| Cellular Component | GO:0062023 | collagen-containing extracellular matrix | HDA | J:265604 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:171712 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:171712 | |||||||||
| Cellular Component | GO:0031410 | cytoplasmic vesicle | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005576 | extracellular region | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005615 | extracellular space | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005615 | extracellular space | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005771 | multivesicular body | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005771 | multivesicular body | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005791 | rough endoplasmic reticulum | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005791 | rough endoplasmic reticulum | IBA | J:265628 | |||||||||
| Biological Process | GO:0002376 | immune system process | IEA | J:60000 | |||||||||
| Biological Process | GO:0043152 | induction of bacterial agglutination | ISO | J:164563 | |||||||||
| Biological Process | GO:0045087 | innate immune response | IEA | J:60000 | |||||||||
| Biological Process | GO:0048286 | lung alveolus development | ISO | J:164563 | |||||||||
| Biological Process | GO:0052403 | negative regulation by host of symbiont catalytic activity | ISO | J:164563 | |||||||||
| Biological Process | GO:0032703 | negative regulation of interleukin-2 production | ISO | J:155856 | |||||||||
| Biological Process | GO:0050765 | negative regulation of phagocytosis | ISO | J:155856 | |||||||||
| Biological Process | GO:0042130 | negative regulation of T cell proliferation | ISO | J:155856 | |||||||||
| Biological Process | GO:0008228 | opsonization | ISO | J:155856 | |||||||||
| Biological Process | GO:0050766 | positive regulation of phagocytosis | ISO | J:155856 | |||||||||
| Biological Process | GO:0050766 | positive regulation of phagocytosis | IBA | J:265628 | |||||||||
| Biological Process | GO:1905226 | regulation of adhesion of symbiont to host epithelial cell | ISO | J:164563 | |||||||||
| Biological Process | GO:0007585 | respiratory gaseous exchange by respiratory system | IEA | J:60000 | |||||||||
| Biological Process | GO:0043129 | surfactant homeostasis | ISO | J:164563 | |||||||||
| Biological Process | GO:0043129 | surfactant homeostasis | ISO | J:155856 | |||||||||
| Biological Process | GO:0043129 | surfactant homeostasis | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/23/2024 MGI 6.23 |

|
|
|
||