|
Symbol Name ID |
Tead3
TEA domain family member 3 MGI:109241 |
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
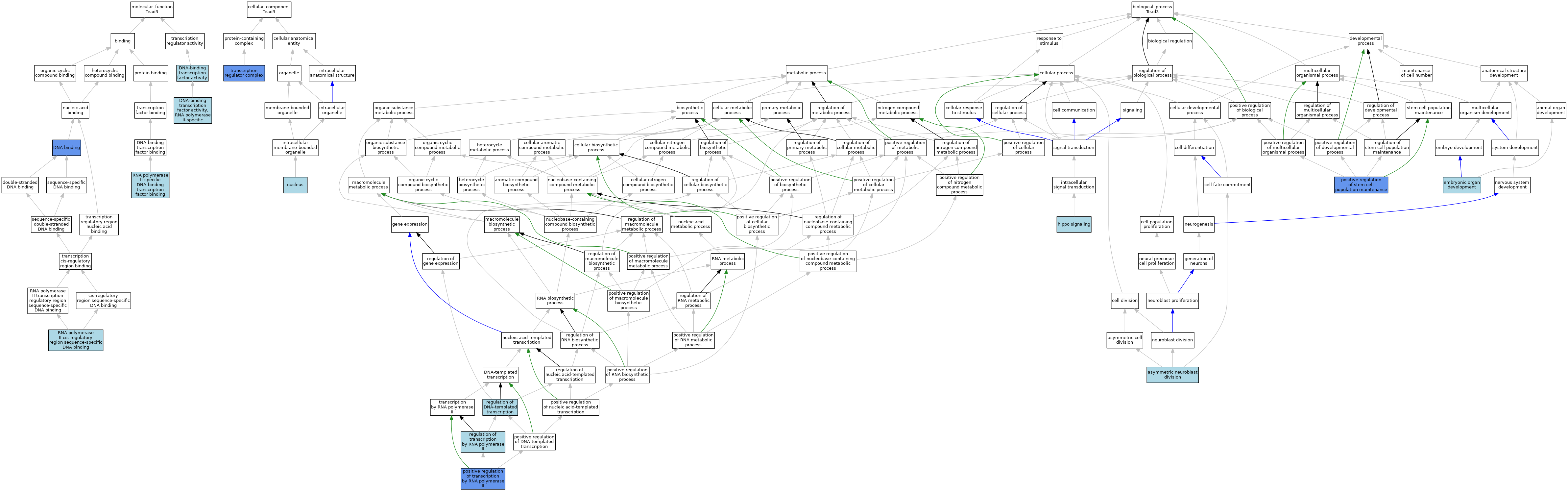
| Molecular Function | GO:0003677 | DNA binding | IDA | J:46262 | |||||||||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0000981 | DNA-binding transcription factor activity, RNA polymerase II-specific | IBA | J:265628 | |||||||||
| Molecular Function | GO:0000978 | RNA polymerase II cis-regulatory region sequence-specific DNA binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0061629 | RNA polymerase II-specific DNA-binding transcription factor binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005667 | transcription regulator complex | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005667 | transcription regulator complex | IGI | J:102951 | |||||||||
| Biological Process | GO:0055059 | asymmetric neuroblast division | ISO | J:164563 | |||||||||
| Biological Process | GO:0048568 | embryonic organ development | IBA | J:265628 | |||||||||
| Biological Process | GO:0035329 | hippo signaling | ISO | J:164563 | |||||||||
| Biological Process | GO:0035329 | hippo signaling | IBA | J:265628 | |||||||||
| Biological Process | GO:1902459 | positive regulation of stem cell population maintenance | IGI | J:160813 | |||||||||
| Biological Process | GO:1902459 | positive regulation of stem cell population maintenance | IGI | J:160813 | |||||||||
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | IDA | J:46262 | |||||||||
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | IGI | J:102951 | |||||||||
| Biological Process | GO:0006355 | regulation of DNA-templated transcription | IEA | J:72247 | |||||||||
| Biological Process | GO:0006357 | regulation of transcription by RNA polymerase II | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/23/2024 MGI 6.23 |

|
|
|
||