|
Symbol Name ID |
Tfam
transcription factor A, mitochondrial MGI:107810 |
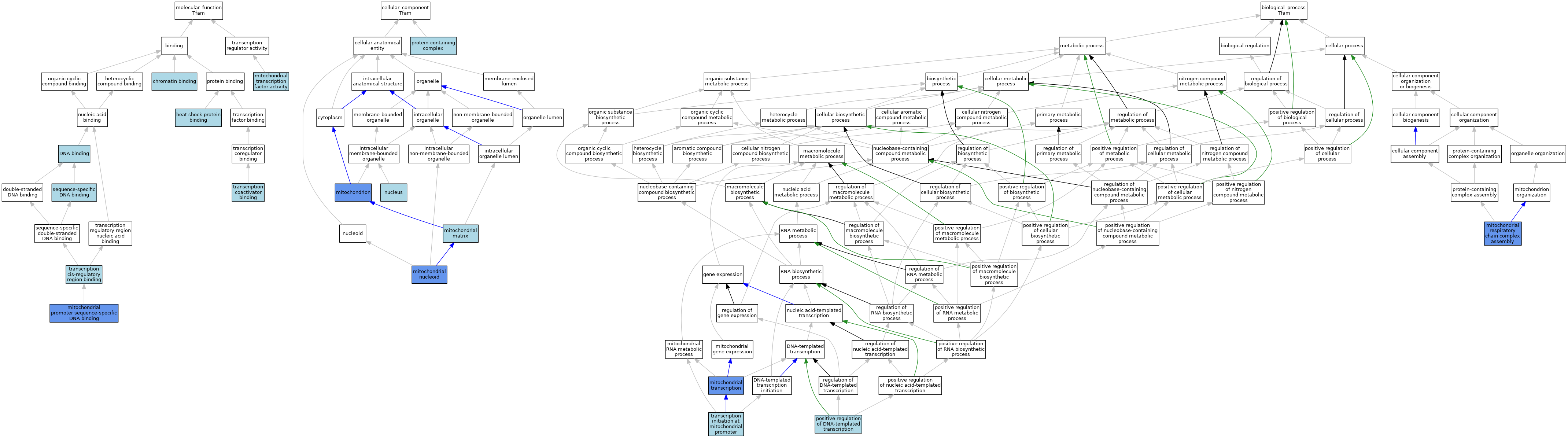
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0003682 | chromatin binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0003677 | DNA binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0031072 | heat shock protein binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0001018 | mitochondrial promoter sequence-specific DNA binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0001018 | mitochondrial promoter sequence-specific DNA binding | IDA | J:256447 | |||||||||
| Molecular Function | GO:0034246 | mitochondrial transcription factor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0043565 | sequence-specific DNA binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0000976 | transcription cis-regulatory region binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0001223 | transcription coactivator binding | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005759 | mitochondrial matrix | ISO | J:164563 | |||||||||
| Cellular Component | GO:0042645 | mitochondrial nucleoid | ISO | J:164563 | |||||||||
| Cellular Component | GO:0042645 | mitochondrial nucleoid | IDA | J:213151 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IDA | J:256447 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:151002 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | IDA | J:33733 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005739 | mitochondrion | HDA | J:86816 | |||||||||
| Cellular Component | GO:0005634 | nucleus | IEA | J:60000 | |||||||||
| Cellular Component | GO:0032991 | protein-containing complex | ISO | J:164563 | |||||||||
| Biological Process | GO:0033108 | mitochondrial respiratory chain complex assembly | IMP | J:125471 | |||||||||
| Biological Process | GO:0006390 | mitochondrial transcription | ISO | J:164563 | |||||||||
| Biological Process | GO:0006390 | mitochondrial transcription | IDA | J:256447 | |||||||||
| Biological Process | GO:0045893 | positive regulation of DNA-templated transcription | ISO | J:164563 | |||||||||
| Biological Process | GO:0006391 | transcription initiation at mitochondrial promoter | ISO | J:164563 | |||||||||
| Biological Process | GO:0006391 | transcription initiation at mitochondrial promoter | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||