|
Symbol Name ID |
Ptpn3
protein tyrosine phosphatase, non-receptor type 3 MGI:105307 |
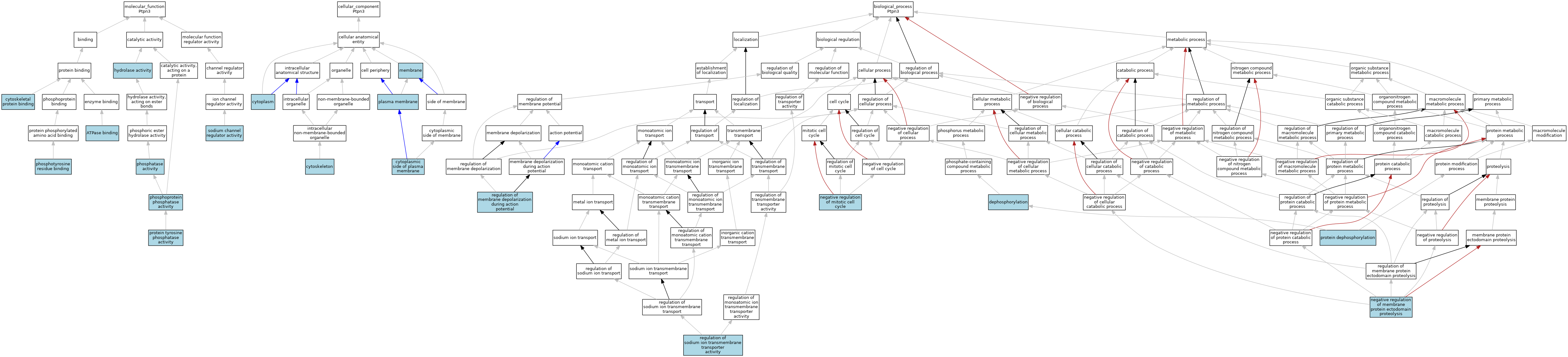
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0051117 | ATPase binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0008092 | cytoskeletal protein binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016787 | hydrolase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016791 | phosphatase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0004721 | phosphoprotein phosphatase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0001784 | phosphotyrosine residue binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004725 | protein tyrosine phosphatase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004725 | protein tyrosine phosphatase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0017080 | sodium channel regulator activity | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IBA | J:265628 | |||||||||
| Cellular Component | GO:0009898 | cytoplasmic side of plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0009898 | cytoplasmic side of plasma membrane | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005856 | cytoskeleton | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005856 | cytoskeleton | IEA | J:72247 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IEA | J:60000 | |||||||||
| Biological Process | GO:0016311 | dephosphorylation | IEA | J:72247 | |||||||||
| Biological Process | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis | ISO | J:164563 | |||||||||
| Biological Process | GO:0045930 | negative regulation of mitotic cell cycle | ISO | J:164563 | |||||||||
| Biological Process | GO:0006470 | protein dephosphorylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0098902 | regulation of membrane depolarization during action potential | ISO | J:164563 | |||||||||
| Biological Process | GO:2000649 | regulation of sodium ion transmembrane transporter activity | ISO | J:164563 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/23/2024 MGI 6.23 |

|
|
|
||