|
Symbol Name ID |
Ephb3
Eph receptor B3 MGI:104770 |
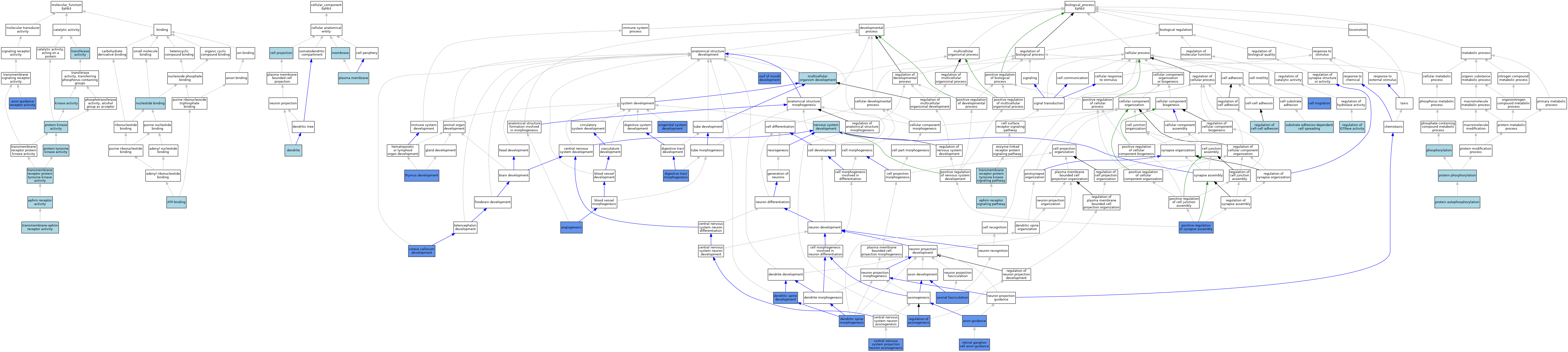
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0008046 | axon guidance receptor activity | IDA | J:71036 | |||||||||
| Molecular Function | GO:0005003 | ephrin receptor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0016301 | kinase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004672 | protein kinase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0004713 | protein tyrosine kinase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004713 | protein tyrosine kinase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004714 | transmembrane receptor protein tyrosine kinase activity | IEA | J:72245 | |||||||||
| Molecular Function | GO:0005005 | transmembrane-ephrin receptor activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0005005 | transmembrane-ephrin receptor activity | TAS | J:71036 | |||||||||
| Cellular Component | GO:0042995 | cell projection | IEA | J:60000 | |||||||||
| Cellular Component | GO:0030425 | dendrite | IBA | J:265628 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:72247 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | TAS | J:71036 | |||||||||
| Biological Process | GO:0001525 | angiogenesis | IMP | J:53085 | |||||||||
| Biological Process | GO:0007411 | axon guidance | IMP | J:37095 | |||||||||
| Biological Process | GO:0007411 | axon guidance | IMP | J:62565 | |||||||||
| Biological Process | GO:0007411 | axon guidance | IBA | J:265628 | |||||||||
| Biological Process | GO:0007411 | axon guidance | IDA | J:71036 | |||||||||
| Biological Process | GO:0007413 | axonal fasciculation | IMP | J:37095 | |||||||||
| Biological Process | GO:0016477 | cell migration | IMP | J:79724 | |||||||||
| Biological Process | GO:0016477 | cell migration | ISO | J:164563 | |||||||||
| Biological Process | GO:0021952 | central nervous system projection neuron axonogenesis | IDA | J:85573 | |||||||||
| Biological Process | GO:0022038 | corpus callosum development | IMP | J:37095 | |||||||||
| Biological Process | GO:0060996 | dendritic spine development | IMP | J:87071 | |||||||||
| Biological Process | GO:0060997 | dendritic spine morphogenesis | IMP | J:87071 | |||||||||
| Biological Process | GO:0048546 | digestive tract morphogenesis | IMP | J:79724 | |||||||||
| Biological Process | GO:0048013 | ephrin receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0048013 | ephrin receptor signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0007275 | multicellular organism development | IEA | J:60000 | |||||||||
| Biological Process | GO:0007399 | nervous system development | IEA | J:60000 | |||||||||
| Biological Process | GO:0016310 | phosphorylation | IEA | J:60000 | |||||||||
| Biological Process | GO:0051965 | positive regulation of synapse assembly | IMP | J:87071 | |||||||||
| Biological Process | GO:0046777 | protein autophosphorylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0006468 | protein phosphorylation | IBA | J:265628 | |||||||||
| Biological Process | GO:0050770 | regulation of axonogenesis | IDA | J:125703 | |||||||||
| Biological Process | GO:0022407 | regulation of cell-cell adhesion | ISO | J:164563 | |||||||||
| Biological Process | GO:0043087 | regulation of GTPase activity | ISO | J:164563 | |||||||||
| Biological Process | GO:0031290 | retinal ganglion cell axon guidance | IDA | J:85573 | |||||||||
| Biological Process | GO:0060021 | roof of mouth development | IMP | J:37095 | |||||||||
| Biological Process | GO:0034446 | substrate adhesion-dependent cell spreading | ISO | J:164563 | |||||||||
| Biological Process | GO:0048538 | thymus development | IMP | J:153788 | |||||||||
| Biological Process | GO:0007169 | transmembrane receptor protein tyrosine kinase signaling pathway | IEA | J:72247 | |||||||||
| Biological Process | GO:0001655 | urogenital system development | IMP | J:91491 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/30/2024 MGI 6.23 |

|
|
|
||