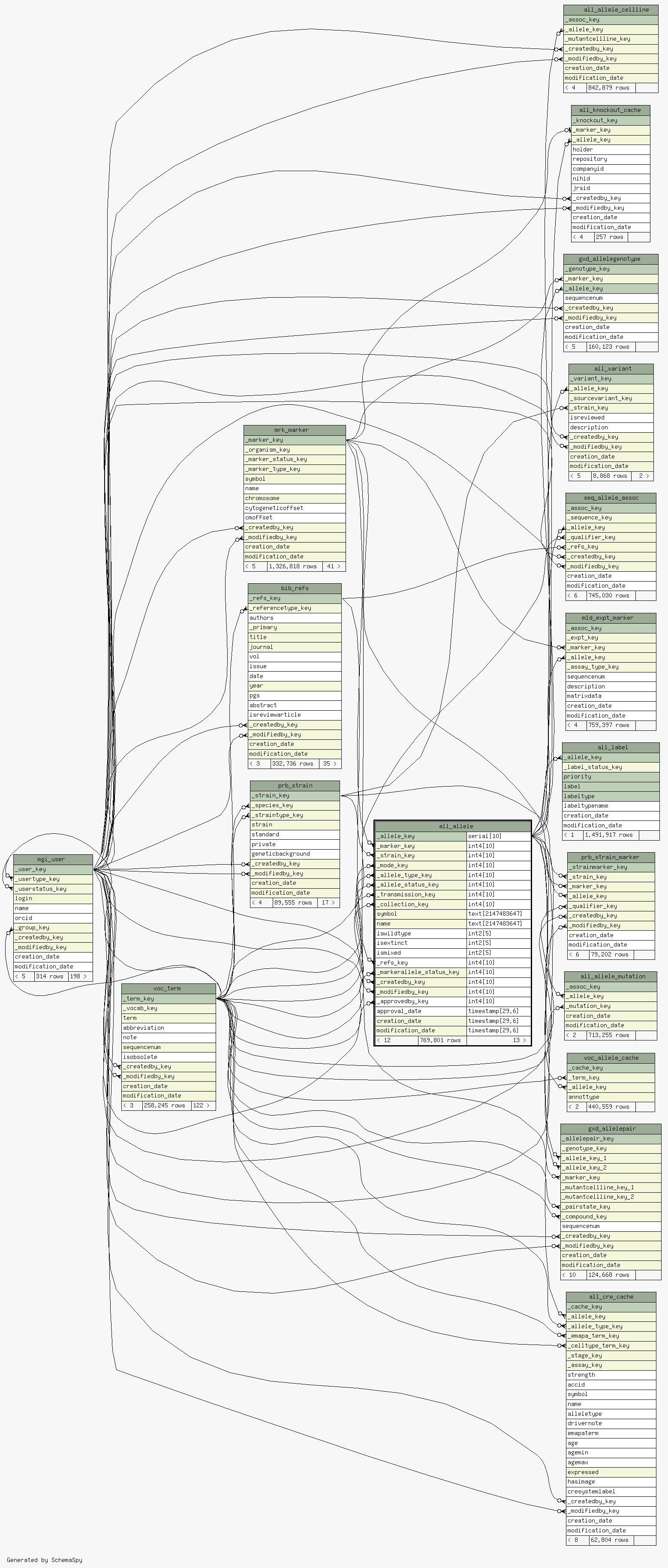
| Table pub.mgd.all_allele A record in this table represents one Allele. An Allele is any of the alternative forms of a gene occupying a given locus; any one of several mutational forms of a gene. In MGI, allelic variants are associated with phenotypes. See ALL_Marker_Assoc. A transgenic Allele will contain a Strain of Origin, and a new Strain can then be bred from this trangene. This new Strain will be associated with the transgenic allele (see PRB_Strain, PRB_Strain_Marker). For example, see TR11000/MMRRC/UC-Davis. For example: MGI:4847055/Tg(Foxi2-EGFP)HX217Gsat, strain of origin: FVB/NTac. New Strain: MGI:5003868/STOCK Tg(Foxi2-EGFP)HX217Gsat/Mmucd associated with MGI:5003868/Tg(Foxi2-EGFP)HX217Gsat via PRB_Strain/PRB_Strain_Marker.
|
Generated by SchemaSpy |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Table contained 808,628 rows at Tue Feb 17 06:11 EST 2026 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Indexes:
| Column(s) | Type | Sort | Constraint Name |
|---|---|---|---|
| _allele_key | Primary key, Used to cluster data | Asc | all_allele_pkey |
| _allele_status_key | Performance | Asc | all_allele_idx_allele_status_key |
| _allele_type_key | Performance | Asc | all_allele_idx_allele_type_key |
| _marker_key | Performance | Asc | all_allele_idx_clustered |
| _collection_key | Performance | Asc | all_allele_idx_collection_key |
| _createdby_key | Performance | Asc | all_allele_idx_createdby_key |
| creation_date | Performance | Asc | all_allele_idx_creation_date |
| _markerallele_status_key | Performance | Asc | all_allele_idx_markerallele_status_key |
| _mode_key | Performance | Asc | all_allele_idx_mode_key |
| modification_date | Performance | Asc | all_allele_idx_modification_date |
| _modifiedby_key | Performance | Asc | all_allele_idx_modifiedby_key |
| name | Performance | Asc | all_allele_idx_name |
| _strain_key | Performance | Asc | all_allele_idx_strain_key |
| symbol | Performance | Asc | all_allele_idx_symbol |
| _transmission_key | Performance | Asc | all_allele_idx_transmission_key |

|
