How do I find genotyping polymorphisms between two strains?
How do I find genotyping polymorphisms between two strains?
| Overview | ||
| You can use the Mouse SNP Query Form to search for mouse single nucleotide polymorphisms that are different between two strains. Use this form to search for SNPs by associated gene, strain or strain comparison or genome coordinates. | ||
| Accessing the Mouse SNP Query Form | ||
|
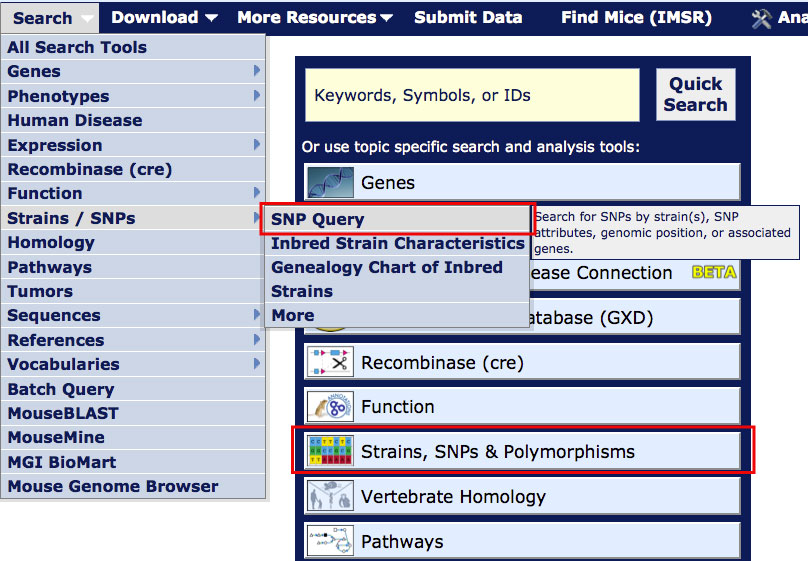
On most MGI pages, you can access the Mouse SNP Query Form
from the Search menu as shown in the image at right.
For this tutorial, open the Mouse SNP Query Form in a new window. Scroll down this page for further instructions. |
|
|
| Example 1. Finding SNPs that are different between A/J and BALB/cByJ | ||
|
|
|
|
|
|