This help document answers the following questions:
What is a BioMart?
BioMart is a query-oriented data management and integration system that performs like a batch query. The system can be applied to a single or to multiple databases. It provides fast, powerful, and complex queries to an individual database, federations (connections) with other databases, and ongoing enhancements. BioMart can be used to convert one or more data sources (flat files or a relational databases) into data marts that are accessible either with a web browser or with Perl, Java and webservice APIs (Application Program Interfaces).
BioMart is a joint development of the Ontario Institute for Cancer Research (OICR) and the European Bioinformatics Institute (EBI).
Top
What are some of the advantages of using MGI BioMart?
BioMart provides an efficient solution to issues such as:
- having to learn a different interface for each software tool that you use
- needing more than one data resource to refine searches
- desiring to integrate your data with other datasets/databases
- spending time cutting and pasting results between websites
- requiring a custom SQL, a web service or an API to accomplish your tasks.
BioMart provides one generic interface, one resource, with no need for a custom SQL/web service/API. You can import data, reiterate and refine queries on one page, and integrate your data with other datasets that MGI BioMart hosts (e.g. Ensembl, VEGA, RGD) and export results, all without cutting and pasting. You can also combine filters and attributes to fully annotate your dataset with a remote one, all without a custom SQL, web service, or API.
Top
What databases are available through the MGI BioMart?
MGI BioMart provides access to 3 databases, composed of datasets.
Databases currently available for access are named:
- MGI Biomart
- Ensembl Mart 57
- VEGA 56
Top
What datasets are available in the MGI BioMart?
MGI Biomart datasets:
- Genes & Genome Features
- Gene Expression Data (GXD).
A dataset is a collection of data tables that follow a given naming convention. Each column in the table represents a variable. Each row corresponds to a member of the dataset. Examples are
- Mus musculus genes from the NCBI m38 mouse assembly (Ensembl and VEGA)
- All public markers in MGI (genes, genome features, transgenes, QTLs, and so on)
- All public alleles in MGI that have been associated to a marker
- Gene expression data (e.g., assay type, developmental stage, anatomical structure, level of expression, pattern).
MGI BioMart filters - restrict a query by selecting:
- Genome feature, location, allele, and ortholog for approximately 30,000 genes in the MGI database
- Gene, stage and tissue of expression, expression level and assay type.
MGI BioMart attributes - determine the data columns displayed in the query results:
- Genome features, locations, alleles, and orthologs for approximately 30,000 genes in the MGI database
- Gene symbol, assay type, stage and anatomical structure and expression results.
Top
How do I use MGI BioMart?
You use MGI BioMart to:
- Query on (or import a list of) genes or genome features or gene expression data, filter it (apply query restrictions), and determine the output sort order.
- Refine your query by adding/subtracting filter.
- Change how the query results display by adding/subtracting attributes.
- Query a different database.
- Integrate MGI data with data from the other database.
- Export the results to a file that you can keep or send to another mart.
To do this:
Click and then select options from the right and left panes of the user interface:
- Select a database to query against.
- (Optional) Apply Filters (e.g. click Choose file to import a list of gene symbols or IDs or to paste them in the box; click to select a Genome Location).
- Select Attributes (data to be returned) (e.g. MGI ID, Feature Name, etc.).
- (Optional) Get an idea of the size of the data returned vs. dataset queried against (click Count)
- Click Results.
- (Optional) Adjust the output (e.g. View more/less data, Export all results to the desired file type and format, check Unique results only).
- (Optional) Refine results and requery, (e.g. add different
filters or attributes, reorder result columns, change output format).
- (Optional) Add another dataset to the initial query.
See also Are there additional tips for using MGI BioMart?
Top
Can I tell in advance the size of the dataset that a query returns?
Yes.
- Click Count (before clicking Results).
- Two numbers appear: the number of items input or found versus the number of items in the selected database.
Top
What is the function of filters?
MGI BioMart uses the concept of filters for narrowing or expanding a query. Initially, you may wish to query the entire set of genes and genome features in MGI, but you can also query a smaller, more limited set (for example, by importing a list of IDs or symbols or selecting a specific region of a chromosome) using the filters. Below are expanded images of all MGI BioMart Genes & Genome Features and Gene Expression Data filter categories.
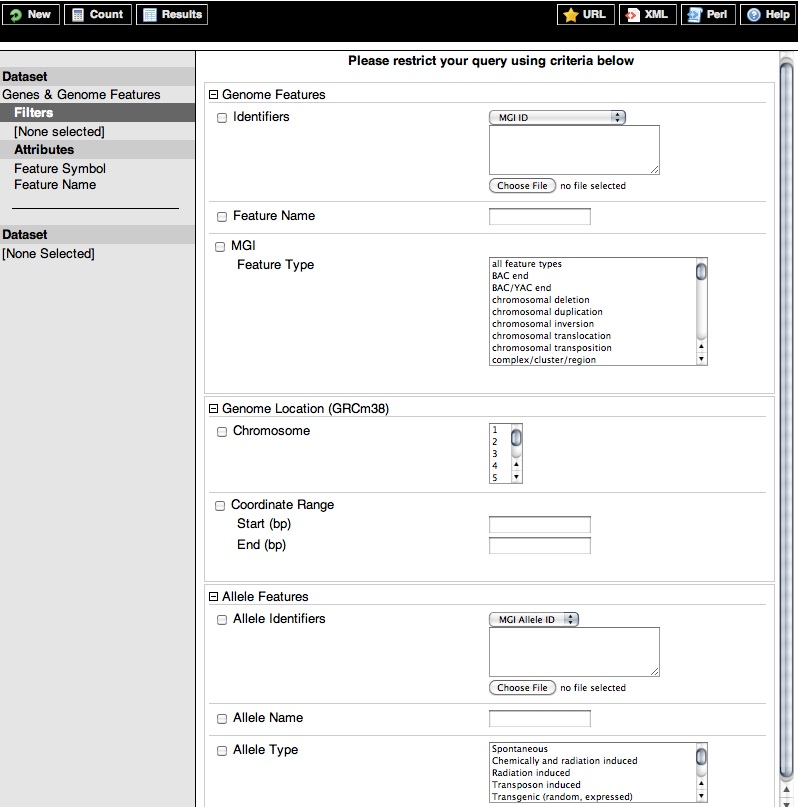
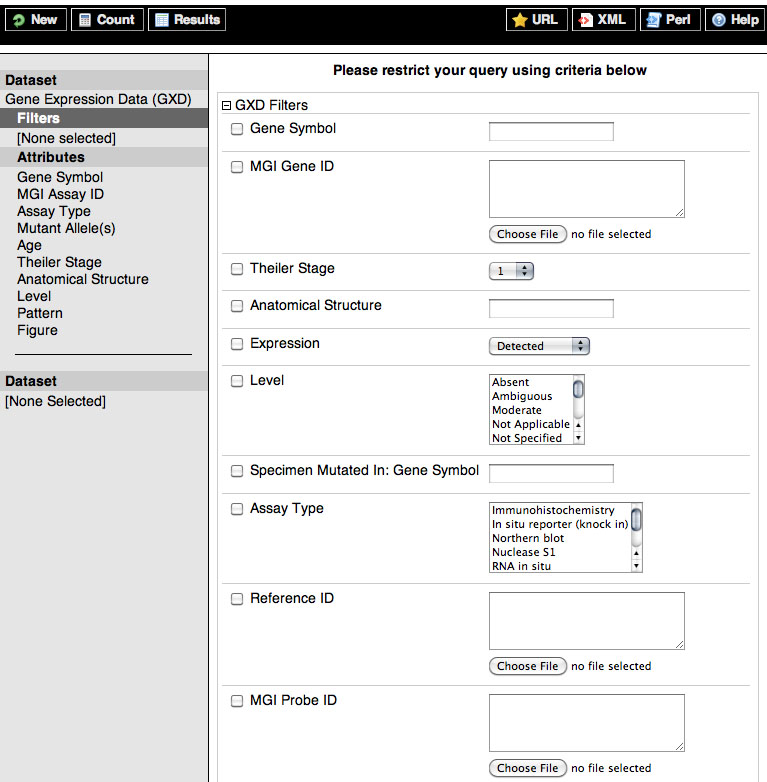
Top
What is the function of attributes?
BioMart identifies the dataset information to add to your query results as attributes. Below are expanded images of MGI BioMart Genes & Genome Features and Gene Expression data (GXD) attribute categories.
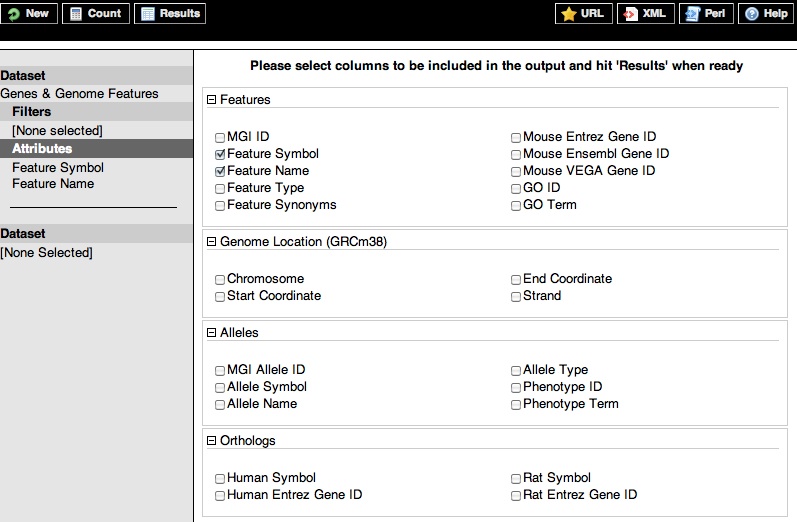
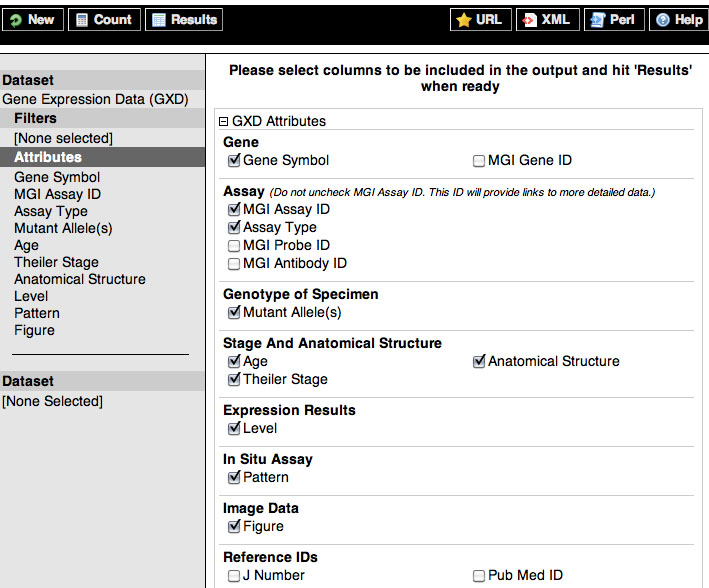
Below are the attribute names to use when querying for Genes & Genome Features data followed by examples of what MGI BioMart returns.
| | Attribute Name | Examples |
| Features |
|---|
| MGI ID | MGI:87904 |
| Feature Symbol | Actb |
| Feature Name | actin, beta |
| Feature Type | protein coding gene |
| Feature Synonyms | Actx |
| Mouse Entrez gene ID | 11461 |
| Mouse Ensembl gene ID | ENSMUSG00000029580 |
| Mouse VEGA gene ID | OTTMUSG00000015100 |
| GO ID (gene ontology identifier) | GO:0006916 |
| GO term | protein kinase binding |
Genome Location
(GRCm38) |
|---|
| Chromosome | 1 thru 19, X, Y, XY, MT, UN |
| Start Coordinate (bp) | 143665420 |
| End Coordinate (bp) | 143668404 |
| strand | + or - |
| Alleles |
|---|
| MGI allele ID | MGI:2180089 |
| Allele Symbol | Actbtm1bhr |
| Allele Name | targeted mutation 1, Richard R. Behringer |
| Allele Type | Targeted (knock-out) |
| Phenotype ID | MP:0003984 |
| Phenotype Term | embryonic growth retardation |
| Orthologs |
|---|
| Human Symbol | Actb |
| Human Entrez Gene ID | 81822 |
| Rat Symbol | ACTB |
| Rat Entrez Gene ID | 60 |
Below are the attribute names to use when querying for gene expression data followed by examples of what MGI BioMart returns.
| GXD Attributes | Attribute Name | Examples |
| Gene | Gene Symbol | Shh |
|---|
| MGI Gene ID | MGI:98297 |
| Assay | MGI Assay ID | MGI:1275014 |
|---|
| Assay Type | RNA in situ |
| MGI Probe ID | MGI:1194645 |
| MGI Antibody ID | MGI:3053439 |
| Genotype of Specimen | Mutant Allele(s) | Shhtm2Amc |
|---|
| Stage And Anatomical Structure | Age | E18.5 |
|---|
| Theiler Stage | TS26 |
| Anatomical Structure | Cerebellum |
| Expression Results | Level | Trace |
|---|
| In Situ Assay | Pattern | Regionally restricted |
|---|
| Reference IDs | J Number | J:108509 |
|---|
| PubMed ID | 16571630 |
Top
How can I use BioMart to search for specific genome feature types?
- The image below depicts a search for all non-coding RNA genes between 11000000 and 31000000 base pairs on Chr. 7.
- The Count for the returned items is 15 out of 78,757 in the database entries searched.
- As you select Attributes for output, they appear in the left-hand frame (in the image, they are Feature Symbol/Name/Type, Chromosome, Start/End Coordinate). These selections determine the content of any columns appearing in your results.
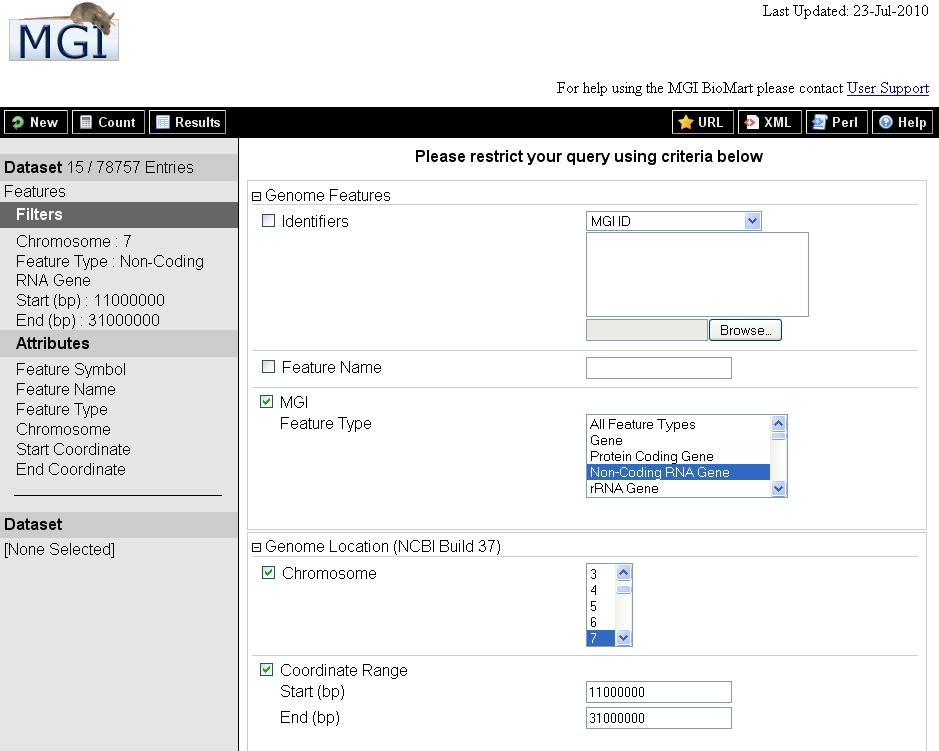
The image below depicts the results of this search.
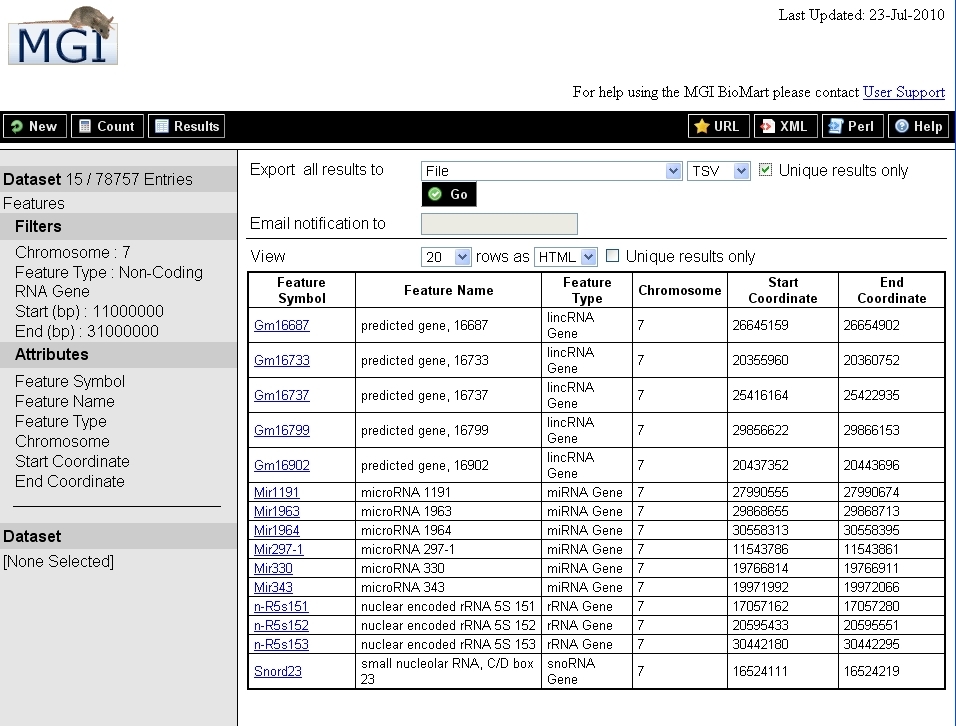
Top
How can I use BioMart to search for gene expression data?
As an example, here is how you can find In situ reporter (knock-in) assays that show expression in diencephalon:
- CHOOSE DATABASE MGI BioMart.
- CHOOSE DATASET Gene Expression Data (GXD).
- Click on Filters in the left hand column to restrict your query.
- Check the box in front of Anatomical Structure and enter diencephalon in its field.
- Check Expression and use its default setting: Detected.
- Check Assay Type and select in situ reporter (knock-in).
- Click the Count button to see the number of expected results. For example: 262 records out of a total of 1181425 (262/1181425).
- Click on Attributes in the left hand column to select the results.
- For this example, just use the default settings.
- Click the Results button.
- If you view your results as HTML, the Figures and MGI Assay IDs link to detail web pages in the MGI Database.
The image below depicts a portion of the results of this search.
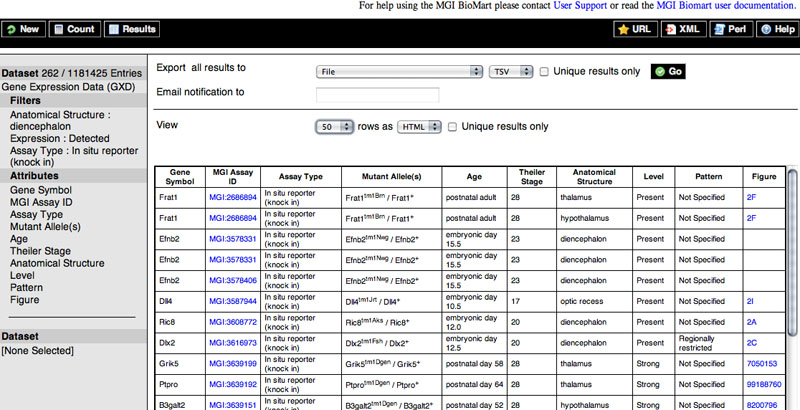
Top
My query returned no results. Why?
- Imported files must contain a single type of identifier (feature symbol or specific ID).
- Imported files (genome or allele features or IDs) must be saved with the .txt extension.
- The proper identifier must appear in the pull-down list. If you leave the default (MGI ID) but are importing a list of feature symbols, there will be no results.
Top
Are there data that MGI BioMart does not currently search for/find?
Yes. The MGI BioMart does not yet contain all MGI data. For example, for the Genes & Genome Features dataset, it does not recognize secondary marker IDs. Secondary IDs are neither counted (e.g. when you click Count) nor do they appear when you click Results.
MGI gene detail pages list secondary IDs in the Other accession IDs section at the bottom. For example, the Kit gene detail page lists MGD-MRK-11588, MGD-MRK-14609, MGD-MRK-15420, MGD-MRK-1672, MGD-MRK-9743, MGI:3530304, MGI:3530312, MGI:3530319. None of these are returned if/when you include Kit in the Feature Symbol list of an MGI BioMart query.
Top
How can I change the sort order in my query results?
The order in which items appear under Attributes (left panel) is the order of the columns in which the query results appear. As an example, for Genes & Genome Feature results, by default, Feature Symbol and Feature Name are the first two columns. To change the order:
- Click Attributes.
- Click to open attribute lists in the right panel (e.g. Features, Genome Location, Alleles, Orthologs).
- Select an item (e.g. MGI ID, Feature Type, etc.) from an attribute list. When you do so, the item appears beneath Attributes in the left-hand panel.
- Deselect any attribute that you do not want to appear as a column in your results.
- Continue selecting/deselecting attributes until they appear in the desired column order (on the left).
- Click Results again.
- Repeat until satisfied with the result.
Top
What can I do on a result page?
- By default, the first 10 results appear in hyper-linked HTML format. To alter the number and format, select from the drop-down lists; e.g. if the result Count is 17 results, select 20 from the View list.
- Select CSV or TSV to see results as comma- or tab-separated values.
- Click Unique results only to remove any redundancy in a given data model. Note, however, that this may affect the time it takes to retrieve your results.
- Once you are satisfied with a result set, choose from the actions at the top of the page and click Go button. If your query is particularly intensive and the server times out, use one of the Web file options.
- You can incorporate a second dataset into the query by clicking on the second Dataset node on the left and following the same steps. Linking of the datasets and merging of data into the final result set is handled automatically.
- To begin a new query, click New (on the toolbar).
- Click the XML button on the toolbar for a summary of your current query in BioMart Query XML format. This is useful for accessing BioMart via the mart service web services interface. Similarly, click the Perl button to generate a BioMart perl API script to run the same query.
Top
Yes. Be aware that:
- MGI BioMart supports only files with a .txt extension. Therefore, be sure to save any files that you wish to import into BioMart with that extension (e.g. lists of symbols or IDs).
- For the Genes & Genome Features dataset, the default identifier for input is MGI ID. If you wish to filter by Feature Symbol or an ID other than MGI and you do not change the default, the query will fail.
- All imported items must be of the same type (e.g. all Mouse VEGA Gene IDs or all Feature Symbols).
- MGI BioMart does not find allele names that include a comma. You must replace the comma with a % (percent sign) to get results.
MGI BioMart does allow you to:
- Choose an additional dataset (for comparison) and select filters and attributes for it.
- Click Count before you click Results to determine the size of the returned dataset. The number appears beside Dataset at the top of the right-hand panel. As an example, selecting miRNA as a feature type returns 625/78757 which means that there are 625 miRNAs among the 78,757 entries in the MGI BioMart database.
- Add additional parameters to a query after it completes (i.e., rerun it after selecting more columns for the output.
- Rearrange the order in which columns appear in your results. The column order in the results is the order in which you select items under Attributes. Note that Feature Symbol and Feature Type are the default Attribute values and therefore appear as the first and second columns unless you deselect them. You can always re-add either term after making additional choices.
- Change the number of results to view (from 10 to 20, 50, 100, 150, 200 or all results) once they appear.
- Build more/less complex queries and rerun them without starting over (by adding or subtracting attributes).
- Export results to a file, to a .gz (compressed file), or to a compressed web file with an email notification.
- Format any results in web (HTML), CSV (comma-separated value), TSV (tab-separated value) or Excel (.xls) files.
Top
Where can I find additional BioMart information?
The BioMart Project website has additional information.
- To access the application, click MartView.
- Additional tabs provide news and documentation.
Top
 Using the MGI BioMart Datasets
Using the MGI BioMart Datasets
 Using the MGI BioMart Datasets
Using the MGI BioMart Datasets


