|
Symbol Name ID |
Yes1
YES proto-oncogene 1, Src family tyrosine kinase MGI:99147 |
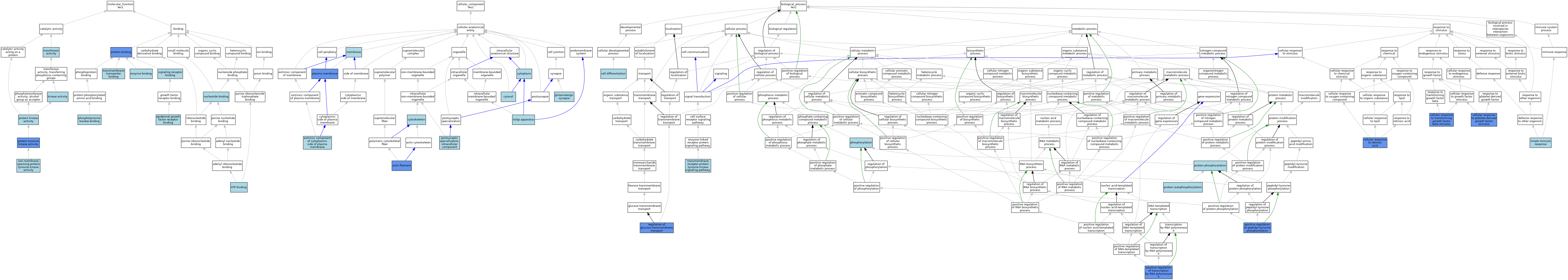
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0019899 | enzyme binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0005154 | epidermal growth factor receptor binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0016301 | kinase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004715 | non-membrane spanning protein tyrosine kinase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0001784 | phosphotyrosine residue binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:72781 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:182976 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:72781 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:72781 | |||||||||
| Molecular Function | GO:0004672 | protein kinase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0004713 | protein tyrosine kinase activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0004713 | protein tyrosine kinase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004713 | protein tyrosine kinase activity | IDA | J:236752 | |||||||||
| Molecular Function | GO:0004713 | protein tyrosine kinase activity | TAS | Reactome:R-MMU-9763891 | |||||||||
| Molecular Function | GO:0005102 | signaling receptor binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0044325 | transmembrane transporter binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005884 | actin filament | IDA | J:125001 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005856 | cytoskeleton | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9680646 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9680706 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9682158 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9682182 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9682572 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9763891 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9763892 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9763903 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9764150 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9817994 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-9818009 | |||||||||
| Cellular Component | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane | IBA | J:265628 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005794 | Golgi apparatus | ISO | J:164563 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IDA | J:125001 | |||||||||
| Cellular Component | GO:0099091 | postsynaptic specialization, intracellular component | ISO | J:155856 | |||||||||
| Biological Process | GO:0030154 | cell differentiation | IBA | J:265628 | |||||||||
| Biological Process | GO:0036120 | cellular response to platelet-derived growth factor stimulus | IDA | J:125001 | |||||||||
| Biological Process | GO:0071300 | cellular response to retinoic acid | IDA | J:182976 | |||||||||
| Biological Process | GO:0071560 | cellular response to transforming growth factor beta stimulus | IGI | J:176254 | |||||||||
| Biological Process | GO:0071560 | cellular response to transforming growth factor beta stimulus | IGI | J:176254 | |||||||||
| Biological Process | GO:0045087 | innate immune response | IBA | J:265628 | |||||||||
| Biological Process | GO:0016310 | phosphorylation | IEA | J:60000 | |||||||||
| Biological Process | GO:0050731 | positive regulation of peptidyl-tyrosine phosphorylation | IDA | J:182976 | |||||||||
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | IGI | J:182976 | |||||||||
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | IGI | J:182976 | |||||||||
| Biological Process | GO:0045944 | positive regulation of transcription by RNA polymerase II | IGI | J:182976 | |||||||||
| Biological Process | GO:0046777 | protein autophosphorylation | ISO | J:155856 | |||||||||
| Biological Process | GO:0006468 | protein phosphorylation | IEA | J:72247 | |||||||||
| Biological Process | GO:0010827 | regulation of glucose transmembrane transport | IDA | J:72781 | |||||||||
| Biological Process | GO:0007169 | transmembrane receptor protein tyrosine kinase signaling pathway | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/09/2024 MGI 6.23 |

|
|
|
||