|
Symbol Name ID |
Grk3
G protein-coupled receptor kinase 3 MGI:87941 |
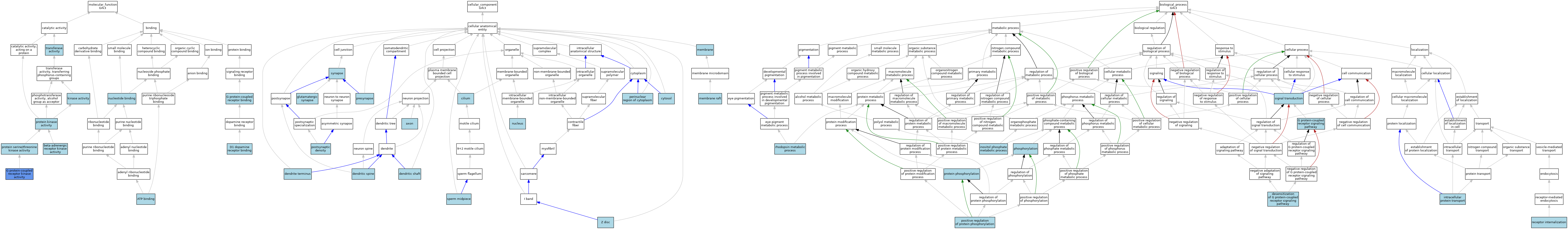
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0047696 | beta-adrenergic receptor kinase activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0031748 | D1 dopamine receptor binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0001664 | G protein-coupled receptor binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0004703 | G protein-coupled receptor kinase activity | ISO | J:155856 | |||||||||
| Molecular Function | GO:0004703 | G protein-coupled receptor kinase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0004703 | G protein-coupled receptor kinase activity | IDA | J:11373 | |||||||||
| Molecular Function | GO:0016301 | kinase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0000166 | nucleotide binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004672 | protein kinase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0004674 | protein serine/threonine kinase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004674 | protein serine/threonine kinase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Cellular Component | GO:0030424 | axon | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005929 | cilium | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:155856 | |||||||||
| Cellular Component | GO:0044292 | dendrite terminus | ISO | J:155856 | |||||||||
| Cellular Component | GO:0043198 | dendritic shaft | ISO | J:155856 | |||||||||
| Cellular Component | GO:0043197 | dendritic spine | ISO | J:155856 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0016020 | membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0045121 | membrane raft | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005634 | nucleus | ISO | J:155856 | |||||||||
| Cellular Component | GO:0048471 | perinuclear region of cytoplasm | ISO | J:155856 | |||||||||
| Cellular Component | GO:0014069 | postsynaptic density | ISO | J:155856 | |||||||||
| Cellular Component | GO:0098793 | presynapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0097225 | sperm midpiece | ISO | J:155856 | |||||||||
| Cellular Component | GO:0045202 | synapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0030018 | Z disc | ISO | J:155856 | |||||||||
| Biological Process | GO:0002029 | desensitization of G protein-coupled receptor signaling pathway | ISO | J:155856 | |||||||||
| Biological Process | GO:0002029 | desensitization of G protein-coupled receptor signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0007186 | G protein-coupled receptor signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:0043647 | inositol phosphate metabolic process | ISO | J:155856 | |||||||||
| Biological Process | GO:0006886 | intracellular protein transport | ISO | J:155856 | |||||||||
| Biological Process | GO:0016310 | phosphorylation | IEA | J:60000 | |||||||||
| Biological Process | GO:0001934 | positive regulation of protein phosphorylation | ISO | J:155856 | |||||||||
| Biological Process | GO:0006468 | protein phosphorylation | ISO | J:155856 | |||||||||
| Biological Process | GO:0006468 | protein phosphorylation | IBA | J:265628 | |||||||||
| Biological Process | GO:0031623 | receptor internalization | ISO | J:164563 | |||||||||
| Biological Process | GO:0046154 | rhodopsin metabolic process | ISO | J:155856 | |||||||||
| Biological Process | GO:0007165 | signal transduction | IEA | J:72247 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||