|
Symbol Name ID |
Crocc
ciliary rootlet coiled-coil, rootletin MGI:3529431 |
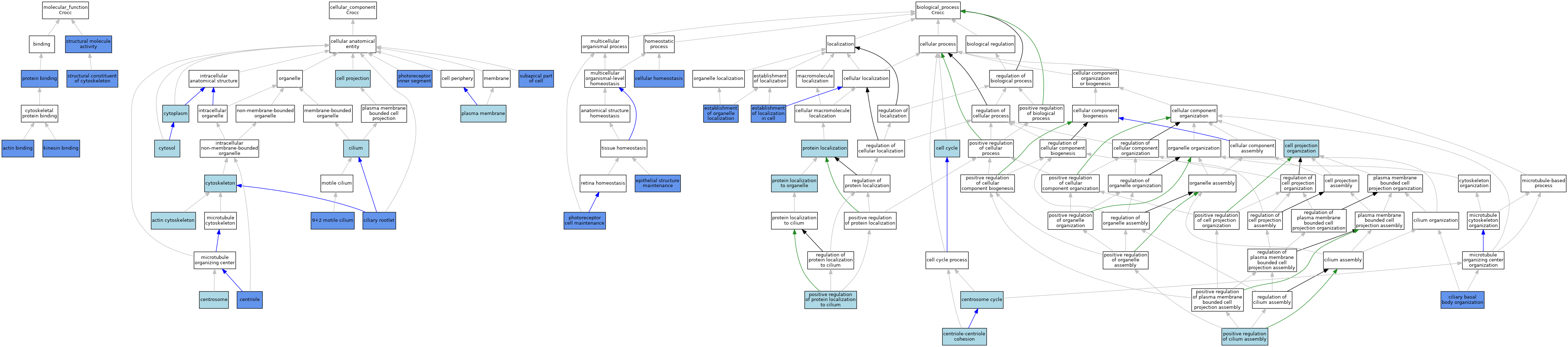
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0003779 | actin binding | IDA | J:98885 | |||||||||
| Molecular Function | GO:0019894 | kinesin binding | IPI | J:102734 | |||||||||
| Molecular Function | GO:0019894 | kinesin binding | IPI | J:102734 | |||||||||
| Molecular Function | GO:0019894 | kinesin binding | IPI | J:102734 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:214070 | |||||||||
| Molecular Function | GO:0005200 | structural constituent of cytoskeleton | IMP | J:98885 | |||||||||
| Molecular Function | GO:0005200 | structural constituent of cytoskeleton | IMP | J:98885 | |||||||||
| Molecular Function | GO:0005198 | structural molecule activity | IDA | J:80161 | |||||||||
| Cellular Component | GO:0097729 | 9+2 motile cilium | IDA | J:267879 | |||||||||
| Cellular Component | GO:0015629 | actin cytoskeleton | ISO | J:164563 | |||||||||
| Cellular Component | GO:0042995 | cell projection | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005814 | centriole | IDA | J:111752 | |||||||||
| Cellular Component | GO:0005814 | centriole | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005814 | centriole | ISO | J:208267 | |||||||||
| Cellular Component | GO:0005813 | centrosome | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005813 | centrosome | IBA | J:265628 | |||||||||
| Cellular Component | GO:0035253 | ciliary rootlet | IDA | J:267879 | |||||||||
| Cellular Component | GO:0035253 | ciliary rootlet | IMP | J:98885 | |||||||||
| Cellular Component | GO:0035253 | ciliary rootlet | IMP | J:193199 | |||||||||
| Cellular Component | GO:0035253 | ciliary rootlet | IDA | J:102734 | |||||||||
| Cellular Component | GO:0035253 | ciliary rootlet | IMP | J:98885 | |||||||||
| Cellular Component | GO:0035253 | ciliary rootlet | IDA | J:193199 | |||||||||
| Cellular Component | GO:0035253 | ciliary rootlet | IDA | J:171542 | |||||||||
| Cellular Component | GO:0005929 | cilium | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005856 | cytoskeleton | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0001917 | photoreceptor inner segment | IDA | J:193199 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0120219 | subapical part of cell | IDA | J:267879 | |||||||||
| Biological Process | GO:0007049 | cell cycle | IEA | J:60000 | |||||||||
| Biological Process | GO:0030030 | cell projection organization | IEA | J:60000 | |||||||||
| Biological Process | GO:0019725 | cellular homeostasis | IMP | J:98885 | |||||||||
| Biological Process | GO:0010457 | centriole-centriole cohesion | ISO | J:164563 | |||||||||
| Biological Process | GO:0007098 | centrosome cycle | ISO | J:164563 | |||||||||
| Biological Process | GO:0007098 | centrosome cycle | IBA | J:265628 | |||||||||
| Biological Process | GO:0032053 | ciliary basal body organization | IMP | J:98885 | |||||||||
| Biological Process | GO:0010669 | epithelial structure maintenance | IMP | J:98885 | |||||||||
| Biological Process | GO:0051649 | establishment of localization in cell | IMP | J:98885 | |||||||||
| Biological Process | GO:0051656 | establishment of organelle localization | IMP | J:98885 | |||||||||
| Biological Process | GO:0045494 | photoreceptor cell maintenance | IMP | J:98885 | |||||||||
| Biological Process | GO:0045724 | positive regulation of cilium assembly | ISO | J:164563 | |||||||||
| Biological Process | GO:1903566 | positive regulation of protein localization to cilium | ISO | J:164563 | |||||||||
| Biological Process | GO:0008104 | protein localization | ISO | J:164563 | |||||||||
| Biological Process | GO:0033365 | protein localization to organelle | ISO | J:164563 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||