|
Symbol Name ID |
Znrf3
zinc and ring finger 3 MGI:3039616 |
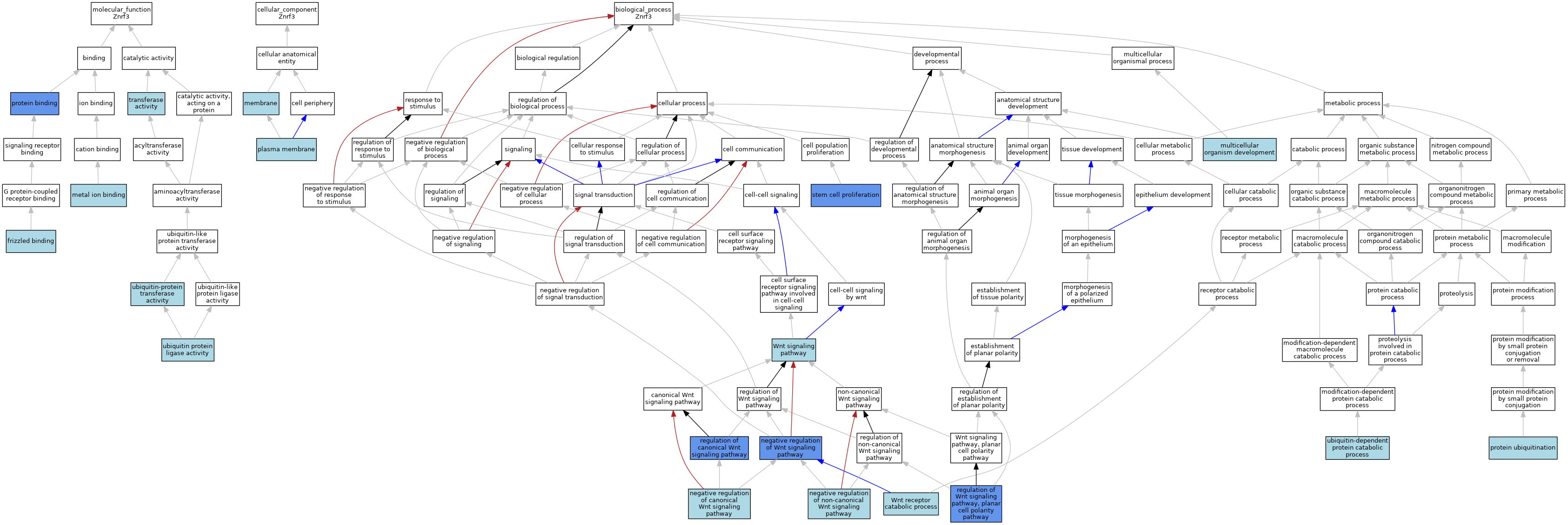
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005109 | frizzled binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0005109 | frizzled binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0046872 | metal ion binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:287053 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:274568 | |||||||||
| Molecular Function | GO:0016740 | transferase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0061630 | ubiquitin protein ligase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0004842 | ubiquitin-protein transferase activity | ISO | J:164563 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Biological Process | GO:0007275 | multicellular organism development | IEA | J:60000 | |||||||||
| Biological Process | GO:0090090 | negative regulation of canonical Wnt signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0090090 | negative regulation of canonical Wnt signaling pathway | IBA | J:265628 | |||||||||
| Biological Process | GO:2000051 | negative regulation of non-canonical Wnt signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0030178 | negative regulation of Wnt signaling pathway | IMP | J:186765 | |||||||||
| Biological Process | GO:0016567 | protein ubiquitination | ISO | J:164563 | |||||||||
| Biological Process | GO:0060828 | regulation of canonical Wnt signaling pathway | IMP | J:184507 | |||||||||
| Biological Process | GO:2000095 | regulation of Wnt signaling pathway, planar cell polarity pathway | IMP | J:184507 | |||||||||
| Biological Process | GO:0072089 | stem cell proliferation | IMP | J:186765 | |||||||||
| Biological Process | GO:0006511 | ubiquitin-dependent protein catabolic process | ISO | J:164563 | |||||||||
| Biological Process | GO:0006511 | ubiquitin-dependent protein catabolic process | IBA | J:265628 | |||||||||
| Biological Process | GO:0038018 | Wnt receptor catabolic process | ISO | J:164563 | |||||||||
| Biological Process | GO:0038018 | Wnt receptor catabolic process | IBA | J:265628 | |||||||||
| Biological Process | GO:0016055 | Wnt signaling pathway | IEA | J:60000 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||