|
Symbol Name ID |
Bloc1s3
biogenesis of lysosomal organelles complex-1, subunit 3 MGI:2678952 |
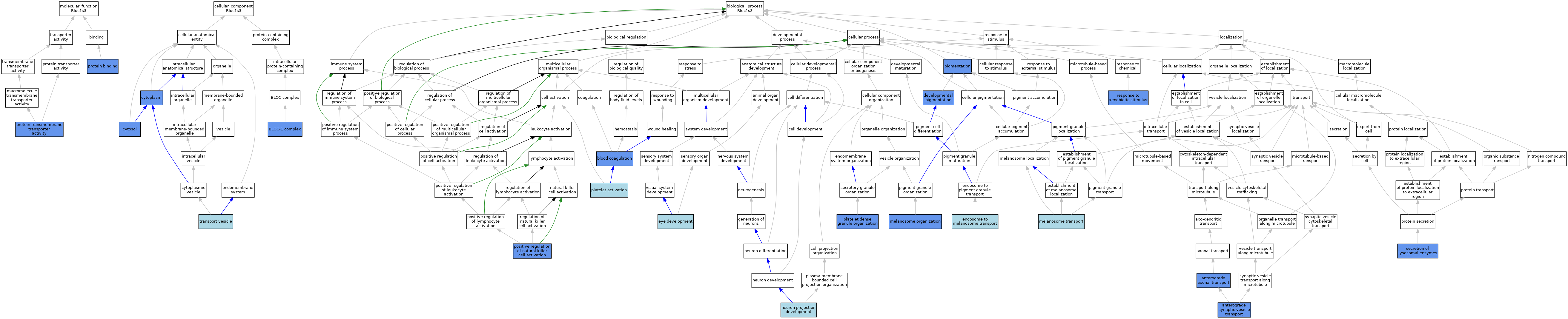
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005515 | protein binding | IPI | J:90887 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:94897 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:90887 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:94897 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:94897 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:90887 | |||||||||
| Molecular Function | GO:0008320 | protein transmembrane transporter activity | IMP | J:114587 | |||||||||
| Cellular Component | GO:0031083 | BLOC-1 complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0031083 | BLOC-1 complex | IDA | J:124026 | |||||||||
| Cellular Component | GO:0031083 | BLOC-1 complex | IBA | J:265628 | |||||||||
| Cellular Component | GO:0031083 | BLOC-1 complex | IDA | J:90887 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IDA | J:94897 | |||||||||
| Cellular Component | GO:0005829 | cytosol | IDA | J:90887 | |||||||||
| Cellular Component | GO:0030133 | transport vesicle | ISO | J:164563 | |||||||||
| Biological Process | GO:0008089 | anterograde axonal transport | IMP | J:187545 | |||||||||
| Biological Process | GO:0048490 | anterograde synaptic vesicle transport | IMP | J:187545 | |||||||||
| Biological Process | GO:0007596 | blood coagulation | IMP | J:94897 | |||||||||
| Biological Process | GO:0048066 | developmental pigmentation | IMP | J:6470 | |||||||||
| Biological Process | GO:0048066 | developmental pigmentation | IMP | J:90887 | |||||||||
| Biological Process | GO:0035646 | endosome to melanosome transport | ISO | J:164563 | |||||||||
| Biological Process | GO:0001654 | eye development | ISO | J:164563 | |||||||||
| Biological Process | GO:0032438 | melanosome organization | NAS | J:90887 | |||||||||
| Biological Process | GO:0032438 | melanosome organization | IMP | J:94897 | |||||||||
| Biological Process | GO:0032402 | melanosome transport | ISO | J:164563 | |||||||||
| Biological Process | GO:0031175 | neuron projection development | NAS | J:188897 | |||||||||
| Biological Process | GO:0043473 | pigmentation | ISO | J:164563 | |||||||||
| Biological Process | GO:0043473 | pigmentation | IMP | J:6801 | |||||||||
| Biological Process | GO:0043473 | pigmentation | IMP | J:7980 | |||||||||
| Biological Process | GO:0043473 | pigmentation | IMP | J:29294 | |||||||||
| Biological Process | GO:0043473 | pigmentation | IMP | J:94897 | |||||||||
| Biological Process | GO:0030168 | platelet activation | ISO | J:164563 | |||||||||
| Biological Process | GO:0060155 | platelet dense granule organization | IMP | J:94897 | |||||||||
| Biological Process | GO:0032816 | positive regulation of natural killer cell activation | IMP | J:6801 | |||||||||
| Biological Process | GO:0009410 | response to xenobiotic stimulus | IMP | J:29294 | |||||||||
| Biological Process | GO:0033299 | secretion of lysosomal enzymes | IMP | J:6801 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/09/2024 MGI 6.23 |

|
|
|
||