|
Symbol Name ID |
Wfikkn1
WAP, FS, Ig, KU, and NTR-containing protein 1 MGI:2670967 |
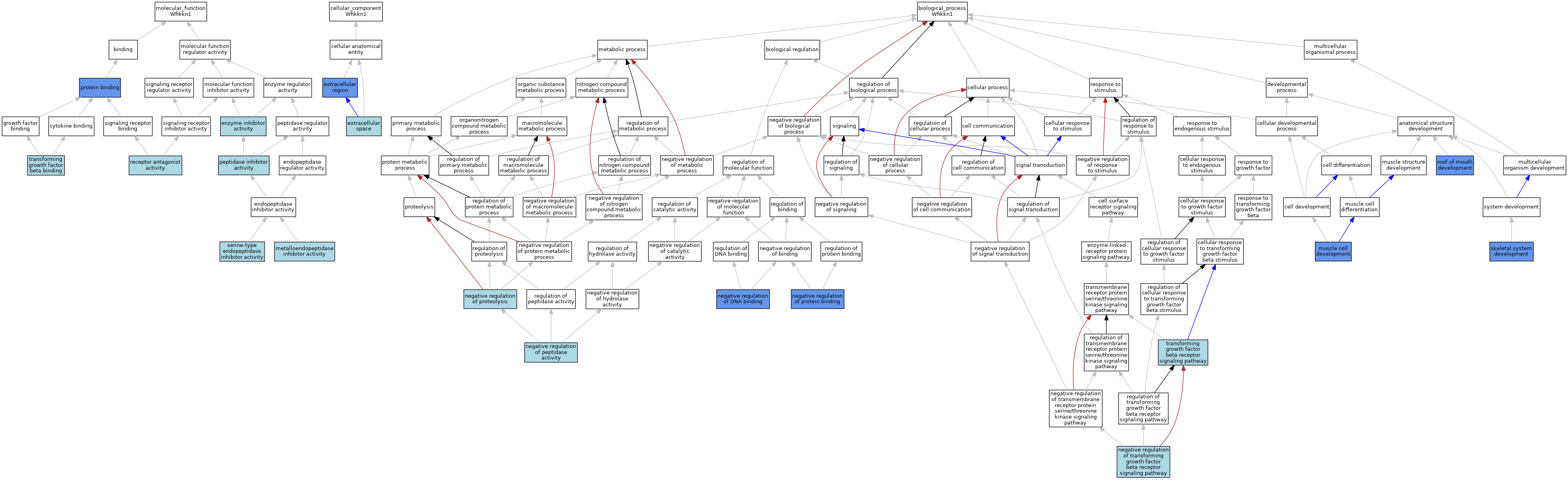
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0004857 | enzyme inhibitor activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0008191 | metalloendopeptidase inhibitor activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0030414 | peptidase inhibitor activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0030414 | peptidase inhibitor activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:201148 | |||||||||
| Molecular Function | GO:0048019 | receptor antagonist activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0048019 | receptor antagonist activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0004867 | serine-type endopeptidase inhibitor activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0004867 | serine-type endopeptidase inhibitor activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0050431 | transforming growth factor beta binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0050431 | transforming growth factor beta binding | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005576 | extracellular region | IDA | J:201148 | |||||||||
| Cellular Component | GO:0005615 | extracellular space | IBA | J:265628 | |||||||||
| Biological Process | GO:0055001 | muscle cell development | IMP | J:201148 | |||||||||
| Biological Process | GO:0043392 | negative regulation of DNA binding | IDA | J:201148 | |||||||||
| Biological Process | GO:0010466 | negative regulation of peptidase activity | IEA | J:60000 | |||||||||
| Biological Process | GO:0032091 | negative regulation of protein binding | IDA | J:201148 | |||||||||
| Biological Process | GO:0045861 | negative regulation of proteolysis | ISO | J:73065 | |||||||||
| Biological Process | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0060021 | roof of mouth development | IGI | J:201148 | |||||||||
| Biological Process | GO:0001501 | skeletal system development | IGI | J:201148 | |||||||||
| Biological Process | GO:0001501 | skeletal system development | IGI | J:201148 | |||||||||
| Biological Process | GO:0007179 | transforming growth factor beta receptor signaling pathway | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||