|
Symbol Name ID |
Cxcl17
C-X-C motif chemokine ligand 17 MGI:2387642 |
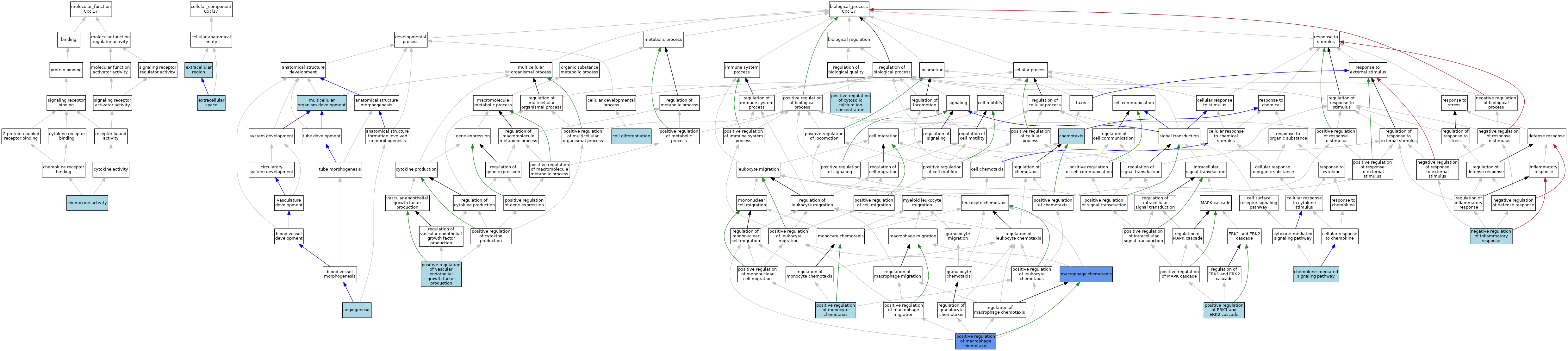
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0008009 | chemokine activity | ISO | J:230699 | |||||||||
| Cellular Component | GO:0005576 | extracellular region | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005615 | extracellular space | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005615 | extracellular space | IBA | J:265628 | |||||||||
| Biological Process | GO:0001525 | angiogenesis | IEA | J:60000 | |||||||||
| Biological Process | GO:0030154 | cell differentiation | IEA | J:60000 | |||||||||
| Biological Process | GO:0070098 | chemokine-mediated signaling pathway | ISO | J:230699 | |||||||||
| Biological Process | GO:0006935 | chemotaxis | IEA | J:60000 | |||||||||
| Biological Process | GO:0048246 | macrophage chemotaxis | IBA | J:265628 | |||||||||
| Biological Process | GO:0048246 | macrophage chemotaxis | IMP | J:230699 | |||||||||
| Biological Process | GO:0007275 | multicellular organism development | IEA | J:60000 | |||||||||
| Biological Process | GO:0050728 | negative regulation of inflammatory response | ISO | J:164563 | |||||||||
| Biological Process | GO:0050728 | negative regulation of inflammatory response | IBA | J:265628 | |||||||||
| Biological Process | GO:0007204 | positive regulation of cytosolic calcium ion concentration | ISO | J:230699 | |||||||||
| Biological Process | GO:0070374 | positive regulation of ERK1 and ERK2 cascade | ISO | J:164563 | |||||||||
| Biological Process | GO:0010759 | positive regulation of macrophage chemotaxis | ISO | J:164563 | |||||||||
| Biological Process | GO:0010759 | positive regulation of macrophage chemotaxis | IMP | J:232054 | |||||||||
| Biological Process | GO:0010759 | positive regulation of macrophage chemotaxis | IBA | J:265628 | |||||||||
| Biological Process | GO:0090026 | positive regulation of monocyte chemotaxis | ISO | J:164563 | |||||||||
| Biological Process | GO:0090026 | positive regulation of monocyte chemotaxis | IBA | J:265628 | |||||||||
| Biological Process | GO:0010575 | positive regulation of vascular endothelial growth factor production | ISO | J:164563 | |||||||||
| Biological Process | GO:0010575 | positive regulation of vascular endothelial growth factor production | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||