|
Symbol Name ID |
Sulf1
sulfatase 1 MGI:2138563 |
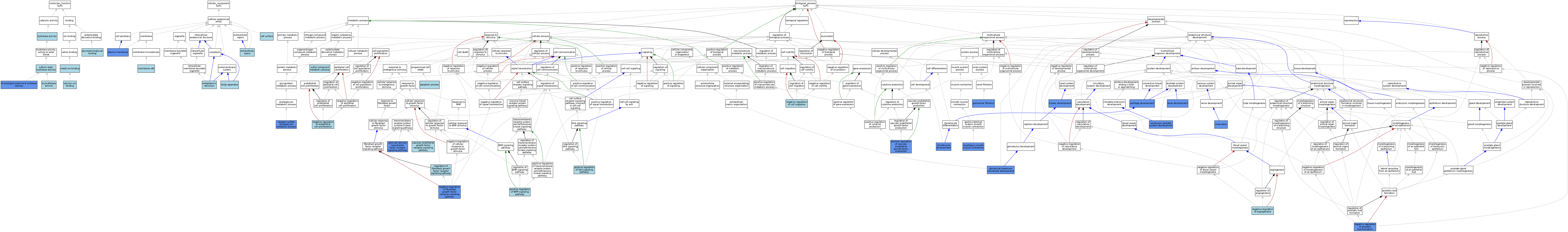
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0004065 | arylsulfatase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0004065 | arylsulfatase activity | ISO | J:80928 | |||||||||
| Molecular Function | GO:0005509 | calcium ion binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0005539 | glycosaminoglycan binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0016787 | hydrolase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0046872 | metal ion binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0008449 | N-acetylglucosamine-6-sulfatase activity | IMP | J:180455 | |||||||||
| Molecular Function | GO:0008449 | N-acetylglucosamine-6-sulfatase activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0008449 | N-acetylglucosamine-6-sulfatase activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0008449 | N-acetylglucosamine-6-sulfatase activity | ISO | J:80928 | |||||||||
| Molecular Function | GO:0008484 | sulfuric ester hydrolase activity | IEA | J:72247 | |||||||||
| Cellular Component | GO:0009986 | cell surface | ISO | J:155856 | |||||||||
| Cellular Component | GO:0009986 | cell surface | ISO | J:164563 | |||||||||
| Cellular Component | GO:0009986 | cell surface | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005783 | endoplasmic reticulum | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005783 | endoplasmic reticulum | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005783 | endoplasmic reticulum | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005615 | extracellular space | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005615 | extracellular space | ISO | J:80928 | |||||||||
| Cellular Component | GO:0005794 | Golgi apparatus | ISO | J:155856 | |||||||||
| Cellular Component | GO:0045121 | membrane raft | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IDA | J:180645 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IBA | J:265628 | |||||||||
| Biological Process | GO:0006915 | apoptotic process | IEA | J:60000 | |||||||||
| Biological Process | GO:0060348 | bone development | IGI | J:129305 | |||||||||
| Biological Process | GO:0051216 | cartilage development | IMP | J:180554 | |||||||||
| Biological Process | GO:0002063 | chondrocyte development | IMP | J:180554 | |||||||||
| Biological Process | GO:0048706 | embryonic skeletal system development | IGI | J:130163 | |||||||||
| Biological Process | GO:0014846 | esophagus smooth muscle contraction | IGI | J:124898 | |||||||||
| Biological Process | GO:0035860 | glial cell-derived neurotrophic factor receptor signaling pathway | IDA | J:124898 | |||||||||
| Biological Process | GO:0032836 | glomerular basement membrane development | IGI | J:180645 | |||||||||
| Biological Process | GO:0003094 | glomerular filtration | IGI | J:180645 | |||||||||
| Biological Process | GO:0030201 | heparan sulfate proteoglycan metabolic process | ISO | J:164563 | |||||||||
| Biological Process | GO:0030201 | heparan sulfate proteoglycan metabolic process | IMP | J:180455 | |||||||||
| Biological Process | GO:0060384 | innervation | IGI | J:124898 | |||||||||
| Biological Process | GO:0001822 | kidney development | IGI | J:129305 | |||||||||
| Biological Process | GO:0016525 | negative regulation of angiogenesis | ISO | J:164563 | |||||||||
| Biological Process | GO:0030336 | negative regulation of cell migration | ISO | J:164563 | |||||||||
| Biological Process | GO:0001937 | negative regulation of endothelial cell proliferation | ISO | J:164563 | |||||||||
| Biological Process | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway | IDA | J:169510 | |||||||||
| Biological Process | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway | IGI | J:180645 | |||||||||
| Biological Process | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway | IGI | J:129305 | |||||||||
| Biological Process | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0060686 | negative regulation of prostatic bud formation | IDA | J:169510 | |||||||||
| Biological Process | GO:0030513 | positive regulation of BMP signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0010575 | positive regulation of vascular endothelial growth factor production | IGI | J:180645 | |||||||||
| Biological Process | GO:0030177 | positive regulation of Wnt signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0040036 | regulation of fibroblast growth factor receptor signaling pathway | ISO | J:164563 | |||||||||
| Biological Process | GO:0006790 | sulfur compound metabolic process | ISO | J:80928 | |||||||||
| Biological Process | GO:0048010 | vascular endothelial growth factor receptor signaling pathway | ISO | J:164563 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/09/2024 MGI 6.23 |

|
|
|
||