|
Symbol Name ID |
Dnaja4
DnaJ heat shock protein family (Hsp40) member A4 MGI:1927638 |
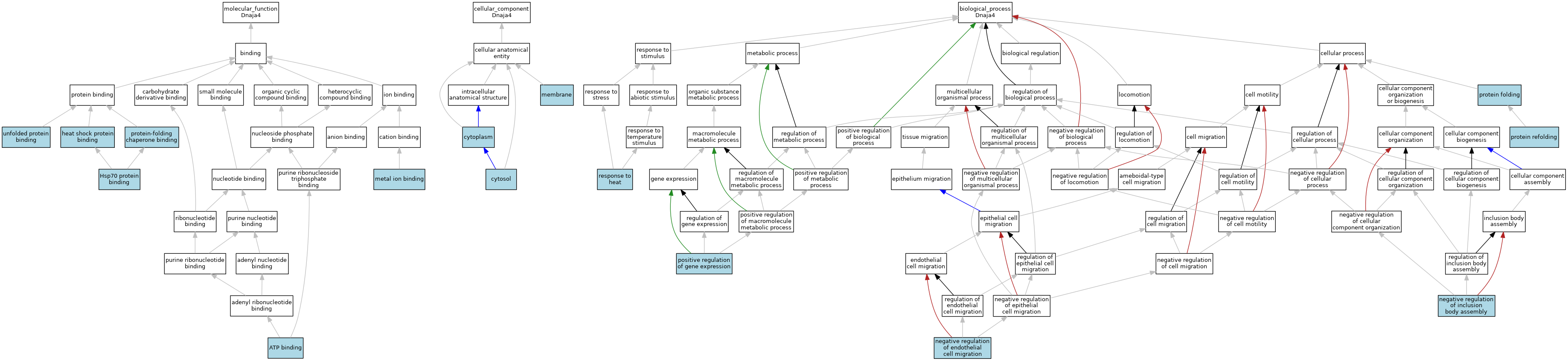
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0031072 | heat shock protein binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0030544 | Hsp70 protein binding | IEA | J:72247 | |||||||||
| Molecular Function | GO:0046872 | metal ion binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0051087 | protein-folding chaperone binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0051087 | protein-folding chaperone binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0051082 | unfolded protein binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005829 | cytosol | IBA | J:265628 | |||||||||
| Cellular Component | GO:0016020 | membrane | ISO | J:164563 | |||||||||
| Biological Process | GO:0010596 | negative regulation of endothelial cell migration | ISO | J:164563 | |||||||||
| Biological Process | GO:0090084 | negative regulation of inclusion body assembly | ISO | J:164563 | |||||||||
| Biological Process | GO:0010628 | positive regulation of gene expression | ISO | J:164563 | |||||||||
| Biological Process | GO:0006457 | protein folding | IEA | J:72247 | |||||||||
| Biological Process | GO:0042026 | protein refolding | ISO | J:164563 | |||||||||
| Biological Process | GO:0042026 | protein refolding | IBA | J:265628 | |||||||||
| Biological Process | GO:0009408 | response to heat | IEA | J:72247 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||