|
Symbol Name ID |
Lin7c
lin-7 homolog C, crumbs cell polarity complex component MGI:1330839 |
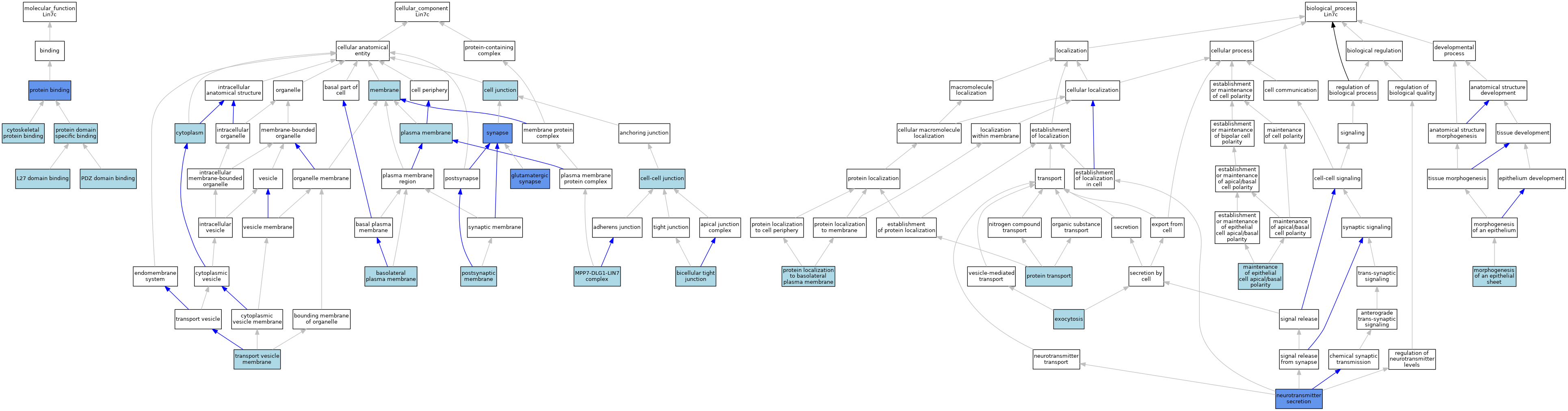
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0008092 | cytoskeletal protein binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0097016 | L27 domain binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0097016 | L27 domain binding | IBA | J:265628 | |||||||||
| Molecular Function | GO:0030165 | PDZ domain binding | ISO | J:155856 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:150861 | |||||||||
| Molecular Function | GO:0019904 | protein domain specific binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0016323 | basolateral plasma membrane | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005923 | bicellular tight junction | IEA | J:60000 | |||||||||
| Cellular Component | GO:0030054 | cell junction | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005911 | cell-cell junction | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005911 | cell-cell junction | ISO | J:155856 | |||||||||
| Cellular Component | GO:0005911 | cell-cell junction | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | ISO | J:164563 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:102649 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:102649 | |||||||||
| Cellular Component | GO:0098978 | glutamatergic synapse | IDA | J:102649 | |||||||||
| Cellular Component | GO:0016020 | membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0097025 | MPP7-DLG1-LIN7 complex | ISO | J:164563 | |||||||||
| Cellular Component | GO:0097025 | MPP7-DLG1-LIN7 complex | IBA | J:265628 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0045211 | postsynaptic membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0045202 | synapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0045202 | synapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0045202 | synapse | IBA | J:265628 | |||||||||
| Cellular Component | GO:0045202 | synapse | IDA | J:102649 | |||||||||
| Cellular Component | GO:0045202 | synapse | IDA | J:102649 | |||||||||
| Cellular Component | GO:0045202 | synapse | IDA | J:102649 | |||||||||
| Cellular Component | GO:0045202 | synapse | IDA | J:102649 | |||||||||
| Cellular Component | GO:0030658 | transport vesicle membrane | TAS | Reactome:R-MMU-9610428 | |||||||||
| Cellular Component | GO:0030658 | transport vesicle membrane | TAS | Reactome:R-MMU-9610641 | |||||||||
| Biological Process | GO:0006887 | exocytosis | IEA | J:60000 | |||||||||
| Biological Process | GO:0045199 | maintenance of epithelial cell apical/basal polarity | IBA | J:265628 | |||||||||
| Biological Process | GO:0002011 | morphogenesis of an epithelial sheet | ISO | J:164563 | |||||||||
| Biological Process | GO:0007269 | neurotransmitter secretion | IBA | J:265628 | |||||||||
| Biological Process | GO:0007269 | neurotransmitter secretion | IGI | J:102649 | |||||||||
| Biological Process | GO:0007269 | neurotransmitter secretion | IGI | J:102649 | |||||||||
| Biological Process | GO:1903361 | protein localization to basolateral plasma membrane | IBA | J:265628 | |||||||||
| Biological Process | GO:0015031 | protein transport | IEA | J:60000 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||