|
Symbol Name ID |
Rasgrp1
RAS guanyl releasing protein 1 MGI:1314635 |
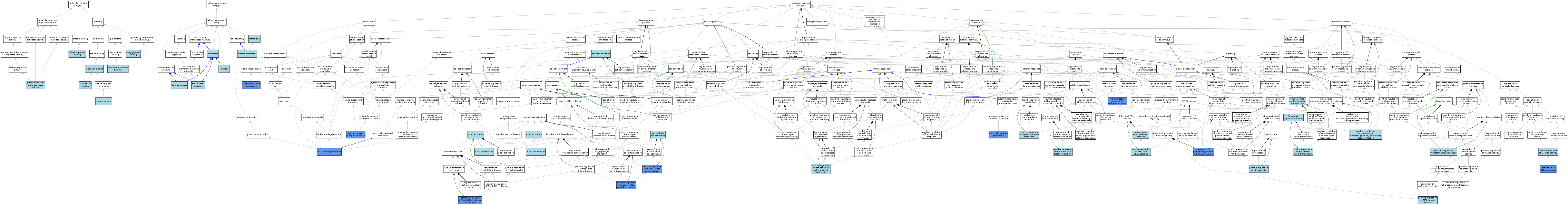
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0005509 | calcium ion binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0019992 | diacylglycerol binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0005085 | guanyl-nucleotide exchange factor activity | ISO | J:164563 | |||||||||
| Molecular Function | GO:0005085 | guanyl-nucleotide exchange factor activity | IBA | J:265628 | |||||||||
| Molecular Function | GO:0042802 | identical protein binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0046872 | metal ion binding | IEA | J:60000 | |||||||||
| Molecular Function | GO:0031210 | phosphatidylcholine binding | ISO | J:164563 | |||||||||
| Molecular Function | GO:0008270 | zinc ion binding | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005829 | cytosol | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-1169494 | |||||||||
| Cellular Component | GO:0005829 | cytosol | TAS | Reactome:R-MMU-2730865 | |||||||||
| Cellular Component | GO:0005783 | endoplasmic reticulum | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005794 | Golgi apparatus | ISO | J:164563 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | ISO | J:164563 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | TAS | Reactome:R-MMU-1169494 | |||||||||
| Cellular Component | GO:0005886 | plasma membrane | IBA | J:265628 | |||||||||
| Biological Process | GO:0090630 | activation of GTPase activity | IMP | J:65115 | |||||||||
| Biological Process | GO:0090630 | activation of GTPase activity | ISO | J:164563 | |||||||||
| Biological Process | GO:0090630 | activation of GTPase activity | IMP | J:125310 | |||||||||
| Biological Process | GO:0042113 | B cell activation | ISO | J:164563 | |||||||||
| Biological Process | GO:0042100 | B cell proliferation | ISO | J:164563 | |||||||||
| Biological Process | GO:0030154 | cell differentiation | IEA | J:60000 | |||||||||
| Biological Process | GO:0002437 | inflammatory response to antigenic stimulus | IMP | J:125310 | |||||||||
| Biological Process | GO:0032762 | mast cell cytokine production | IMP | J:125310 | |||||||||
| Biological Process | GO:0043303 | mast cell degranulation | IMP | J:125310 | |||||||||
| Biological Process | GO:0030101 | natural killer cell activation | ISO | J:164563 | |||||||||
| Biological Process | GO:0070374 | positive regulation of ERK1 and ERK2 cascade | ISO | J:164563 | |||||||||
| Biological Process | GO:0032725 | positive regulation of granulocyte macrophage colony-stimulating factor production | ISO | J:164563 | |||||||||
| Biological Process | GO:0043547 | positive regulation of GTPase activity | ISO | J:164563 | |||||||||
| Biological Process | GO:0043547 | positive regulation of GTPase activity | IBA | J:265628 | |||||||||
| Biological Process | GO:0046330 | positive regulation of JNK cascade | ISO | J:164563 | |||||||||
| Biological Process | GO:0043406 | positive regulation of MAP kinase activity | ISO | J:164563 | |||||||||
| Biological Process | GO:0032816 | positive regulation of natural killer cell activation | IMP | J:179434 | |||||||||
| Biological Process | GO:0032825 | positive regulation of natural killer cell differentiation | IMP | J:179434 | |||||||||
| Biological Process | GO:0045954 | positive regulation of natural killer cell mediated cytotoxicity | ISO | J:164563 | |||||||||
| Biological Process | GO:0001934 | positive regulation of protein phosphorylation | ISO | J:164563 | |||||||||
| Biological Process | GO:0046579 | positive regulation of Ras protein signal transduction | ISO | J:164563 | |||||||||
| Biological Process | GO:0033089 | positive regulation of T cell differentiation in thymus | IMP | J:65115 | |||||||||
| Biological Process | GO:0032760 | positive regulation of tumor necrosis factor production | ISO | J:164563 | |||||||||
| Biological Process | GO:0032729 | positive regulation of type II interferon production | ISO | J:164563 | |||||||||
| Biological Process | GO:0007265 | Ras protein signal transduction | IBA | J:265628 | |||||||||
| Biological Process | GO:0070372 | regulation of ERK1 and ERK2 cascade | ISO | J:164563 | |||||||||
| Biological Process | GO:0014066 | regulation of phosphatidylinositol 3-kinase signaling | IMP | J:125310 | |||||||||
| Biological Process | GO:0032252 | secretory granule localization | IMP | J:125310 | |||||||||
| Biological Process | GO:0007264 | small GTPase mediated signal transduction | IEA | J:72247 | |||||||||
| Biological Process | GO:0042110 | T cell activation | ISO | J:164563 | |||||||||
| Biological Process | GO:0042098 | T cell proliferation | ISO | J:164563 | |||||||||
| Biological Process | GO:0047496 | vesicle transport along microtubule | IMP | J:125310 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/16/2024 MGI 6.23 |

|
|
|
||