|
Symbol Name ID |
Ptprn2
protein tyrosine phosphatase receptor type N polypeptide 2 MGI:107418 |
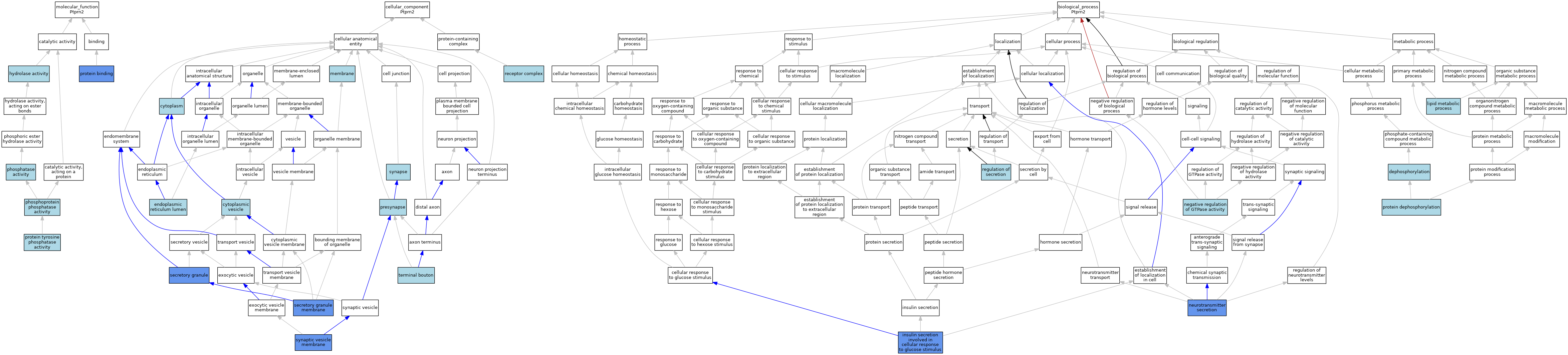
| Category | GO ID | Classification Term | Evidence | Reference | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0016787 | hydrolase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0016791 | phosphatase activity | IEA | J:72247 | |||||||||
| Molecular Function | GO:0004721 | phosphoprotein phosphatase activity | IEA | J:60000 | |||||||||
| Molecular Function | GO:0005515 | protein binding | IPI | J:200322 | |||||||||
| Molecular Function | GO:0004725 | protein tyrosine phosphatase activity | IEA | J:72245 | |||||||||
| Molecular Function | GO:0004725 | protein tyrosine phosphatase activity | IEA | J:72247 | |||||||||
| Cellular Component | GO:0005737 | cytoplasm | ISO | J:155856 | |||||||||
| Cellular Component | GO:0031410 | cytoplasmic vesicle | IEA | J:60000 | |||||||||
| Cellular Component | GO:0005788 | endoplasmic reticulum lumen | ISO | J:155856 | |||||||||
| Cellular Component | GO:0016020 | membrane | IEA | J:60000 | |||||||||
| Cellular Component | GO:0098793 | presynapse | ISO | J:155856 | |||||||||
| Cellular Component | GO:0043235 | receptor complex | ISO | J:194542 | |||||||||
| Cellular Component | GO:0030141 | secretory granule | ISO | J:155856 | |||||||||
| Cellular Component | GO:0030141 | secretory granule | IBA | J:265628 | |||||||||
| Cellular Component | GO:0030141 | secretory granule | IDA | J:100746 | |||||||||
| Cellular Component | GO:0030667 | secretory granule membrane | IDA | J:148933 | |||||||||
| Cellular Component | GO:0045202 | synapse | IBA | J:265628 | |||||||||
| Cellular Component | GO:0030672 | synaptic vesicle membrane | IDA | J:148933 | |||||||||
| Cellular Component | GO:0030672 | synaptic vesicle membrane | ISO | J:155856 | |||||||||
| Cellular Component | GO:0043195 | terminal bouton | ISO | J:155856 | |||||||||
| Biological Process | GO:0016311 | dephosphorylation | IEA | J:72247 | |||||||||
| Biological Process | GO:0035773 | insulin secretion involved in cellular response to glucose stimulus | IMP | J:174841 | |||||||||
| Biological Process | GO:0035773 | insulin secretion involved in cellular response to glucose stimulus | IBA | J:265628 | |||||||||
| Biological Process | GO:0006629 | lipid metabolic process | IEA | J:60000 | |||||||||
| Biological Process | GO:0034260 | negative regulation of GTPase activity | ISO | J:155856 | |||||||||
| Biological Process | GO:0007269 | neurotransmitter secretion | IMP | J:148933 | |||||||||
| Biological Process | GO:0006470 | protein dephosphorylation | IEA | J:72247 | |||||||||
| Biological Process | GO:0051046 | regulation of secretion | IBA | J:265628 | |||||||||
Mouse Genome Database (MGD), Gene Expression Database (GXD), Mouse Models of Human Cancer database (MMHCdb) (formerly Mouse Tumor Biology (MTB)), Gene Ontology (GO) |
||
|
Citing These Resources Funding Information Warranty Disclaimer, Privacy Notice, Licensing, & Copyright Send questions and comments to User Support. |
last database update 04/09/2024 MGI 6.23 |

|
|
|
||