How do I search for all targeted or spontaneous alleles
associated with a gene or marker?
How do I search for all targeted or spontaneous alleles
associated with a gene or marker?
Overview
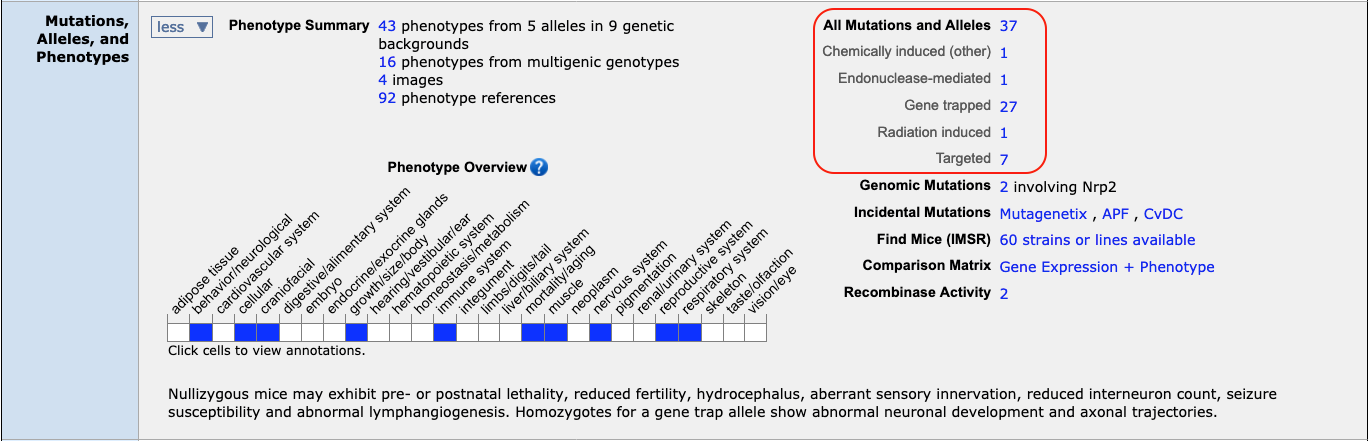
For a single gene search, you can easily retrieve all alleles (or all
transgenic,
all
targeted, all
gene trapped, etc.) by going to the gene detail (ex.
Nrp2) page and clicking on the appropriate
hyperlink in the Mutations, alleles and phenotypes row.
For more customizable options, to search multiple markers simultaneously, or find alleles of any gene associated with a specific phenotype use the Phenotypes, Alleles and Diseases query form.
| Accessing the Phenotypes, Alleles and Disease Models Query Form | ||
|
On most MGI pages, you can access the Phenotypes, Alleles
and Disease Models Query Form from the Search menu as
shown in the image at right. It is also linked on the mini homepage for Phenotypes
& Mutant Alleles.
For this tutorial, open the Phenotypes, Alleles and Diseases query form. in a new window. Scroll down this page for further instructions. |
|
|
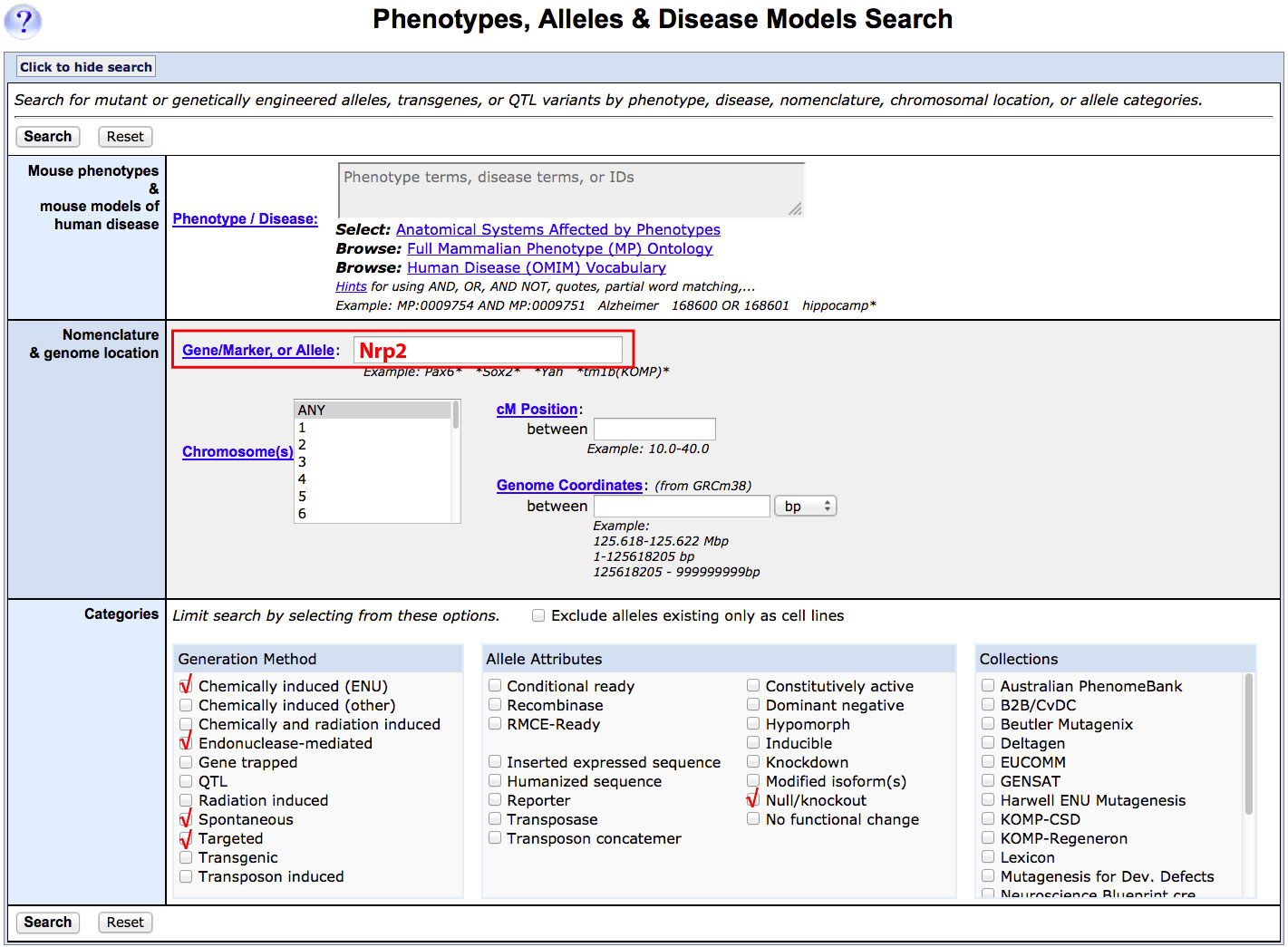
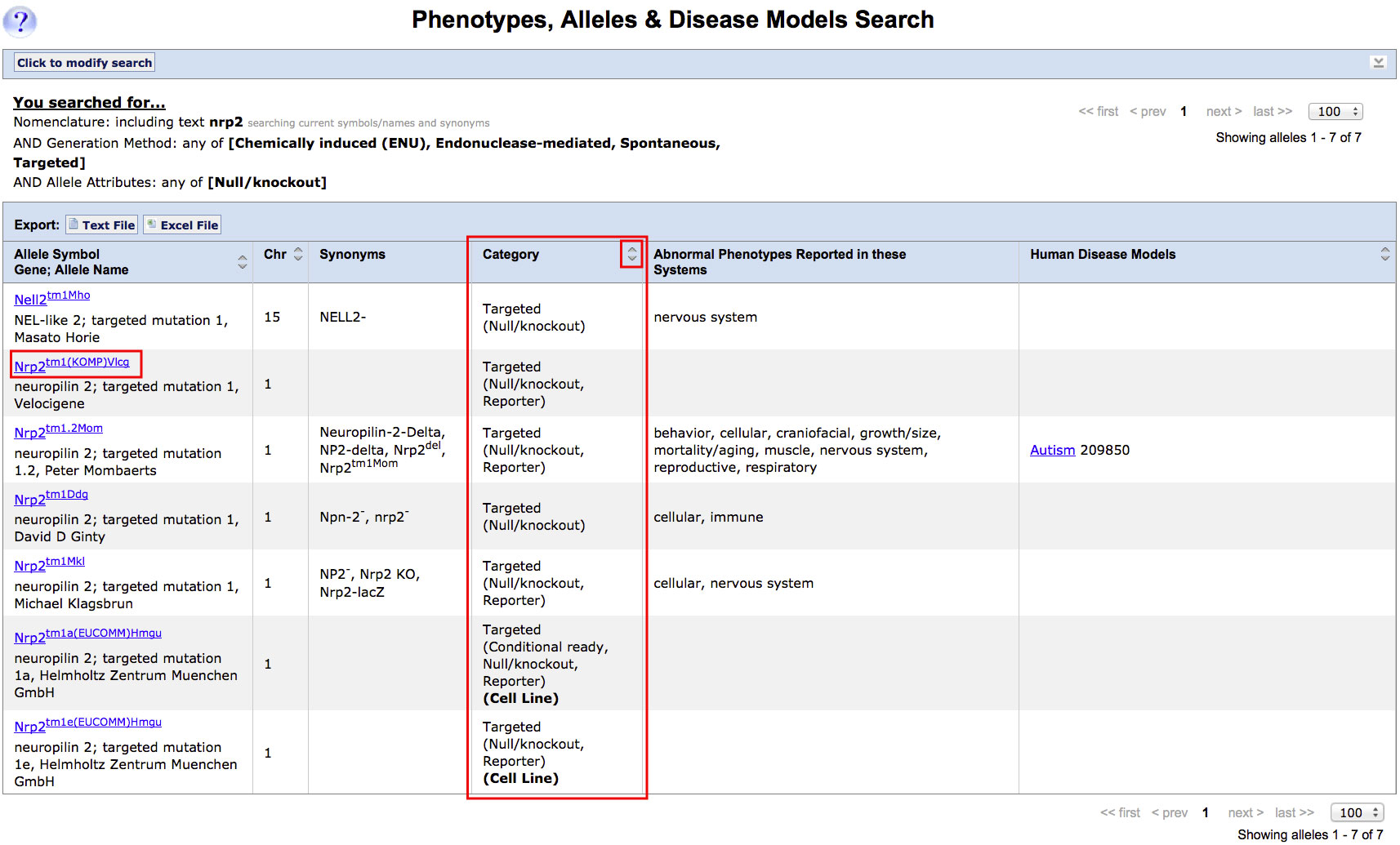
| Example: Finding targeted, spontaneous, ENU-induced or endonuclease mediated mutations in Nrp2 that result in a gene knock-out or null allele. | |||
|
|
||
|
|
||