| Category | ID | Term | Human | Mouse | Rat | Chicken | Zfish | Fly | Worm | Dicty | Dicot | Yeast | Pombe | Ecoli |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
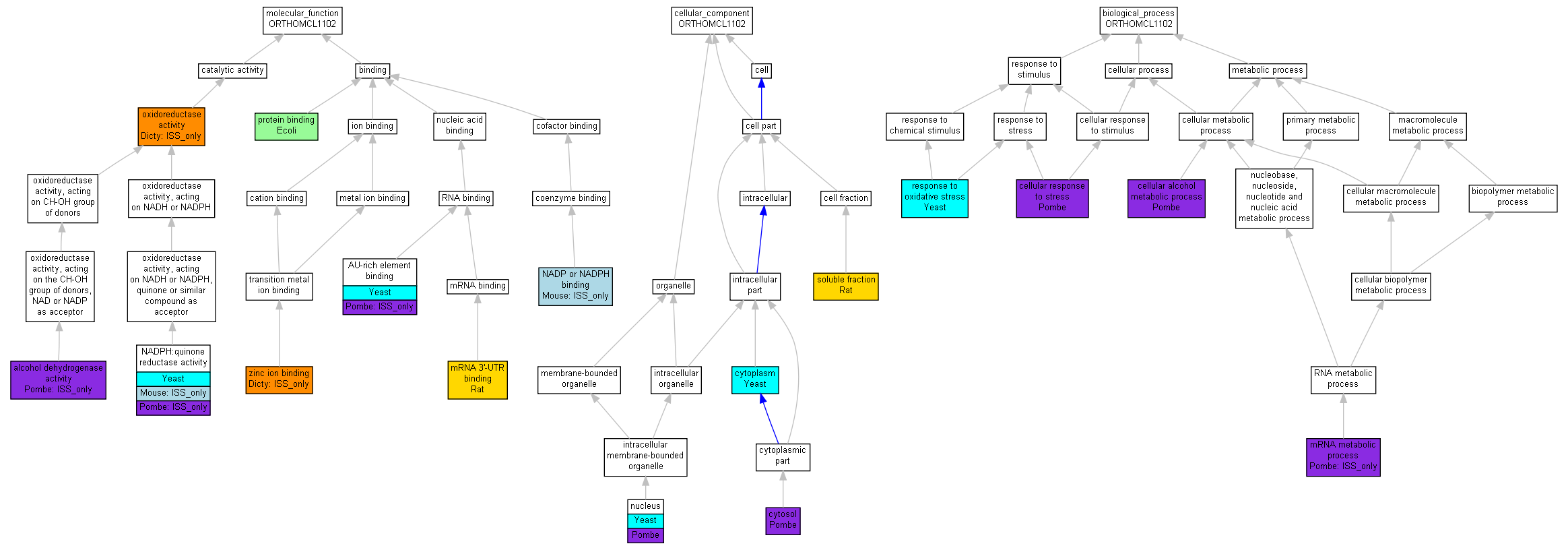
| Molecular Function | GO:0005488 | binding | Human | (Mouse) | Rat | Chicken | Zfish | X | Worm | (Dicty) | Dicot | Yeast | (Pombe) | Ecoli |
| Molecular Function | GO:0003824 | catalytic activity | Human | (Mouse) | Rat | Chicken | Zfish | X | Worm | (Dicty) | Dicot | Yeast | (Pombe) | Ecoli |
| Cellular Component | GO:0005623 | cell | Human | Mouse | Rat | Chicken | Zfish | X | Worm | Dicty | Dicot | Yeast | Pombe | Ecoli |
| Cellular Component | GO:0044464 | cell part | Human | Mouse | Rat | Chicken | Zfish | X | Worm | Dicty | Dicot | Yeast | Pombe | Ecoli |
| Cellular Component | GO:0043226 | organelle | Human | Mouse | Rat | Chicken | Zfish | X | Worm | Dicty | Dicot | Yeast | Pombe | Ecoli |
| Biological Process | GO:0009987 | cellular process | Human | Mouse | Rat | Chicken | Zfish | X | Worm | Dicty | Dicot | Yeast | Pombe | Ecoli |
| Biological Process | GO:0008152 | metabolic process | Human | Mouse | Rat | Chicken | Zfish | X | Worm | Dicty | Dicot | Yeast | Pombe | Ecoli |
| Biological Process | GO:0050896 | response to stimulus | Human | Mouse | Rat | Chicken | Zfish | X | Worm | Dicty | Dicot | Yeast | Pombe | Ecoli |
| Category | ID | Term | DB Object | Evidence | With_From | Organism | Reference |
|---|---|---|---|---|---|---|---|
| Molecular Function | GO:0004022 | alcohol dehydrogenase activity | SPBC1773.06c | ISS | Pfam:PF08240 | Pombe | PMID:17072883 |
| Molecular Function | GO:0017091 | AU-rich element binding | ZTA1 | IDA | Yeast | SGD_REF:S000122743|PMID:17497241 | |
| Molecular Function | GO:0017091 | AU-rich element binding | zta1 | ISS | SGD:S000000250 | Pombe | PMID:17497241 |
| Molecular Function | GO:0003730 | mRNA 3'-UTR binding | Cryz | IDA | Rat | RGD:1601355|PMID:12684230 | |
| Molecular Function | GO:0050661 | NADP or NADPH binding | Cryz | ISO | UniProtKB:O97764 | Mouse | MGI:MGI:87552|PMID:9154917 |
| Molecular Function | GO:0003960 | NADPH:quinone reductase activity | Cryz | ISO | UniProtKB:O97764 | Mouse | MGI:MGI:87552|PMID:9154917 |
| Molecular Function | GO:0003960 | NADPH:quinone reductase activity | ZTA1 | IDA | Yeast | SGD_REF:S000122743|PMID:17497241 | |
| Molecular Function | GO:0003960 | NADPH:quinone reductase activity | zta1 | ISS | SGD:S000000250 | Pombe | PMID:17497241 |
| Molecular Function | GO:0016491 | oxidoreductase activity | DDB_G0271884 | ISS | InterPro:IPR002085 | Dicty | dictyBase_REF:10155 |
| Molecular Function | GO:0016491 | oxidoreductase activity | DDB_G0272440 | ISS | InterPro:IPR002085 | Dicty | dictyBase_REF:10155 |
| Molecular Function | GO:0005515 | protein binding | qor | IPI | UniProtKB:P0A6Y8 | Ecoli | PMID:15690043 |
| Molecular Function | GO:0008270 | zinc ion binding | DDB_G0271884 | ISS | InterPro:IPR002085 | Dicty | dictyBase_REF:10155 |
| Molecular Function | GO:0008270 | zinc ion binding | DDB_G0272440 | ISS | InterPro:IPR002085 | Dicty | dictyBase_REF:10155 |
| Cellular Component | GO:0005737 | cytoplasm | ZTA1 | IDA | Yeast | SGD_REF:S000074185|PMID:14562095 | |
| Cellular Component | GO:0005829 | cytosol | SPBC1773.06c | IDA | Pombe | PMID:16823372 | |
| Cellular Component | GO:0005829 | cytosol | zta1 | IDA | Pombe | PMID:16823372 | |
| Cellular Component | GO:0005634 | nucleus | ZTA1 | IDA | Yeast | SGD_REF:S000074185|PMID:14562095 | |
| Cellular Component | GO:0005634 | nucleus | zta1 | IDA | Pombe | PMID:16823372 | |
| Cellular Component | GO:0005625 | soluble fraction | Cryz | IDA | Rat | RGD:1601355|PMID:12684230 | |
| Biological Process | GO:0006066 | cellular alcohol metabolic process | SPBC1773.06c | IC | GO:0004022 | Pombe | PMID:17072883 |
| Biological Process | GO:0033554 | cellular response to stress | SPBC1773.06c | IEP | GOC:pombekw2GO | Pombe | PMID:12529438 |
| Biological Process | GO:0016071 | mRNA metabolic process | zta1 | ISS | SGD:S000000250 | Pombe | PMID:17497241 |
| Biological Process | GO:0006979 | response to oxidative stress | ZTA1 | IMP | Yeast | SGD_REF:S000122743|PMID:17497241 |
| EXP Inferred from experiment |
| IC Inferred by curator |
| IDA Inferred from direct assay |
| IEA Inferred from electronic annotation |
| IEP Inferred from expression pattern |
| IGC Inferred from genomic context |
| IGI Inferred from genetic interaction |
| IMP Inferred from mutant phenotype |
| IPI Inferred from physical interaction |
| ISA Inferred from sequence alignment |
| ISM Inferred from sequence model |
| ISO Inferred from sequence orthology |
| ISS Inferred from sequence or structural similarity |
| NAS Non-traceable Author Statement |
| ND No biological data available |
| RCA Reviewed computational analysis |
| TAS Traceable author statement |
| NR Not recorded (appears in some legacy annotations) |