Dclre1ctm1Jsek
Targeted Allele Detail
|
|
| Symbol: |
Dclre1ctm1Jsek |
| Name: |
DNA cross-link repair 1C; targeted mutation 1, JoAnn Sekiguchi |
| MGI ID: |
MGI:3843046 |
| Synonyms: |
Artemis-P70, ArtP70 |
| Gene: |
Dclre1c Location: Chr2:3425168-3465167 bp, + strand Genetic Position: Chr2, 1.93 cM
|
| Alliance: |
Dclre1ctm1Jsek page
|
|
|
|
| Allele Type: |
|
Targeted |
| Mutations: |
|
Insertion, Nucleotide substitutions
|
| |
|
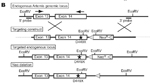
Mutation details: Two point mutations were introduced into exon 14 to substitute a premature stop codon for the aspartate at amino acid position 449 (D449X). This mimics the "Artemis-P70" allele in humans where a 7-nt deletion results in a frameshift at D451 followed by a premature stop codon 10 amino acids downstream. For targeting purposes, a loxP-flanked neomycin resistance cassette was included downstream of exon 14 that was subsequently removed by crossing with cre transgenic mice. Expression of the mutant allele was similar to the endogenous allele as determined by Northern and Q-PCR analysis of splenic RNA from homozygote mice.
(J:147864)
|
|
Generation of the Dclre1ctm1Jsek allele |
|
|
View phenotypes and curated references for all genotypes (concatenated display).
|
|
|
|
|
|
|
| Mouse strains and cell lines
available from the International Mouse Strain Resource
(IMSR) |
| Carrying this Mutation: |
Mouse Strains: 0 strains available
Cell Lines: 0 lines available
|
| Carrying any Dclre1c Mutation: |
30 strains or lines available
|
|
| Original: |
J:147864 Huang Y, et al., Impact of a hypomorphic Artemis disease allele on lymphocyte development, DNA end processing, and genome stability. J Exp Med. 2009 Apr 13;206(4):893-908 |
| All: |
2 reference(s) |
|


